- Title
-
Lhx5 promotes forebrain development and activates transcription of secreted Wnt antagonists
- Authors
- Peng, G., and Westerfield, M.
- Source
- Full text @ Development
|
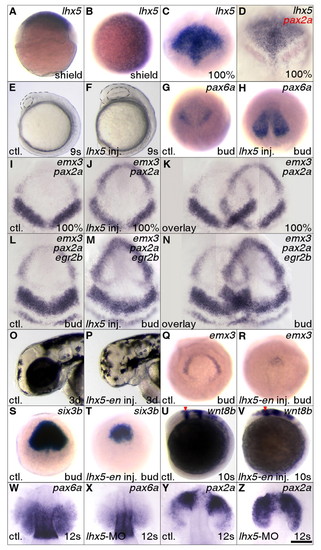
Lhx5 promotes forebrain development. In this and subsequent figures, the probes used for whole-mount in situ hybridization are listed in the upper right corner of each panel. Genotypes or experimental manipulations are indicated in the lower left corners. Developmental stages are indicated in the lower right corners. Unless otherwise noted, gastrula stage embryos are orientated in animal pole view, rostral to the top; post-gastrulation stage embryos in lateral view, rostral to the top and dorsal to the right. ctl, control embryos. (A-D) lhx5 is expressed in rostral regions during embryonic development. (A,B) Dorsal to the right, lateral (A) and animal pole (B) views. (D) A gap can be seen between the lhx5 and pax2a expression domains. (E-N) lhx5 gain of function causes expansion of the forebrain. (E,F) Forebrain boundaries are marked by broken lines. (G,H) Dorsal view, bud stage, pax6a expression. (I-N) Dorsal view, rostral to the top. Embryos were dissected and flat mounted in glycerol after in situ hybridization. Partial overlays of panels were made with PhotoShop (Adobe) and brightness in the overlapped regions was adjusted so that the backgrounds match. (O-V) Inhibition of Lhx5 function compromises forebrain development. (U,V) Red arrowheads mark the forebrain-midbrain boundary. (W-Z) lhx5 morpholino knockdown alters pax6a and pax2a expression. Embryos were dissected and flat mounted in glycerol after in situ hybridization. Dorsal view, rostral to the top. Scale bar in Z: 250 μm for A-H,Q-V; 200 μm for I-K; 167 μm for L-N; 150 μm for O,P; 100 μm for W-Z. EXPRESSION / LABELING:
|
|
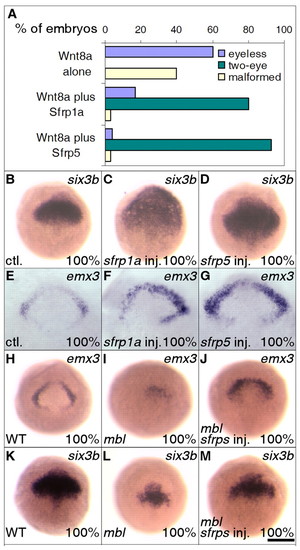
Lhx5 partially rescues forebrain deficiencies induced by ectopic Wnt signaling. (A) lhx5 gain of function partially rescues the eyeless phenotype of wnt8a mRNA-injected embryos. Injected embryos were scored after the Prim-5 stage (24 hours post-fertilization). The percentages of embryos eyeless (blue) or with two eyes (green), or malformed as a result of dorsalization or injection (yellow), were measured. (B-D) lhx5 gain of function partially rescues eye development in mbl (axin1) mutants. Ventral views, rostral to the left. (E-G) lhx5 gain of function partially restores emx3 expression in mbl (axin1) mutants. Genotypes of mutant and partially rescued embryos were verified by RFLP. (H-J) lhx5 gain of function partially restores six3b expression in mbl (axin1) mutants. WT, wild-type embryos. Scale bar in J: 150 μm for B-D; 250 μm for E-J. EXPRESSION / LABELING:
|
|
Sfrps antagonize Wnt signaling and partially restore forebrain marker expression in mbl mutant embryos. (A) Sfrp1a and Sfrp5 rescue eye development in wnt8a mRNA-injected embryos. Injected embryos were scored after 24 hours post-fertilization. (B-D) sfrp1a or sfrp5 gain of function expands the six3b expression domain. (E-G) sfrp1a or sfrp5 gain of function expands the emx3 expression domain. Embryos were dissected and flat mounted in glycerol after in situ hybridization. Animal pole views, rostral to the top. (H-M) Overexpression of Sfrps partially restores emx3 (J) and six3b (M) expression in mbl embryos. Dosage injected was 150 pg of sfrp1a mRNA and 100 pg of sfrp5 mRNA per embryo. Genotypes of mutant and partially rescued embryos were verified by RFLP. WT, wild-type embryos. Scale bar in M: 250 μM for B-M. EXPRESSION / LABELING:
|
|
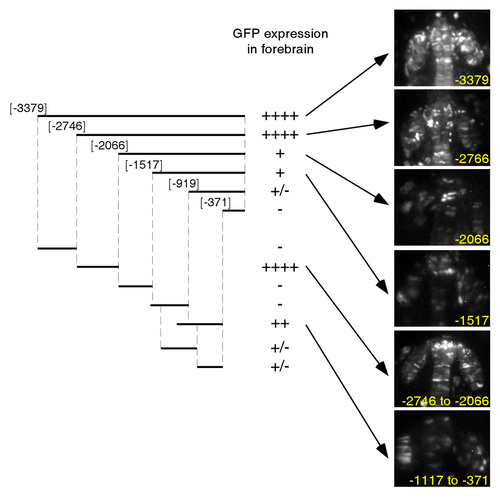
Lhx5 regulates Sfrp1a and Sfrp5 expression. (A-I) Lhx5 regulates Sfrp1a expression. (A-C) Animal pole views, dorsal to the right. (D-F) Dorsal views, bud stage, sfrp1a expression. (G-I) Flat mounts in glycerol; dorsal views, rostral to the top. 2 ng (H) or 4 ng (I) of lhx5 morpholino was injected. (J-M) Lhx5 cell autonomously regulates Sfrp1a expression. Transplanted cells are labeled in red. sfrp1a transcripts are highlighted by white arrowheads and are apparent as blue dots in the narrow cytoplasm around the nuclei of transplanted cells in controls (J, ctl) and lhx5 mRNA-injected cells transplanted into lhx5 morpholino injected hosts (M). sfrp1a transcripts are not detected in lhx5-en injected cells transplanted into control hosts (K). In transplanted lhx5 morpholino-injected cells, sfrp1a expression is reduced (L). (J-L) 90% epiboly stage; (M) one-somite stage. (N) Mapping of sfrp1a upstream enhancers by transient GFP reporter expression in the forebrain. Fragments are numbered using the first nucleotide of the Sfrp1a start codon as the reference. GFP expression in the forebrain was scored at the 3 somite stage and the Prim-5 stage. The number of + signs indicate relative GFP expression levels. (O) Lhx5 binds to the sfrp1a promoter. Input represents 0.2% (2 μl, 2x) or 0.1% (1 μl, 1x) of starting material for each immunoprecipitation reaction. 2 μl (2x) or 1 μl (1x) of purified DNA product from each sample was used in the PCR. ctl., uninjected embryos treated in parallel; inj., lhx5-GFP mRNA-injected embryos. Mock immunoprecipitation with no antibody fails to amplify from either uninjected or injected groups (data not shown). (P-U) Lhx5 regulates Sfrp5 expression. Flat mounts in glycerol, dorsal views, rostral to the top. (R,T,U) 2 ng (R,T) or 4 ng (U) of lhx5 morpholinos was injected. Scale bar in U: 250 μm for A-F; 100 μm for G-I; 8 μm for J-M; 100 μm for P-U. |
|
Sfrp compensates for inhibition of Lhx5 function.(A) Sfrp1a and Sfrp5 rescue eye development in lhx5-en mRNA-injected embryos. The percentages of embryos malformed, as a result of dorsalization or injection (yellow), eyeless (blue) or two eyed (green) were measured. (B-E) Ectopic Wnt signaling results in ectopic axin2 expression. (B,C) Animal pole views; (D,E) lateral views, rostral to the left, dorsal to the top. Genotypes were determined by RFLP. (F-I) Ectopic axin2 expression in lhx5 morpholino-injected embryos. Flat mounts in glycerol. Brain boundaries are marked by broken lines. Green arrows indicate the borders of axin2 labeling. (F,G) Dorsal views, rostral to the top; (H,I) ateral views, rostral to the left, dorsal to the top. Eyes were removed to expose the brain. (J-M) sfrp1a or sfrp5 gain of function reduces ectopic axin2 expression in lhx5-en-injected embryos. Animal pole views, dorsal to the left. Scale bar in M: 250 μm for B,C; 125 μm for D,E; 100 μm for F-I; 250 μm for J-M. EXPRESSION / LABELING:
|
|
Efficacy and sepcificity of lhx5 morpholinos. (A-D) lhx5 morpholino knockdown alters pax6a and pax2a expression patterns. Embryos were dissected and flat mounted in glycerol after in situ hybridization. Dorsal views, rostral to the top. (E-G) lhx5 translation-blocking morpholino reduces lhx5-GFP mRNA translation. GFP fluorescence in the cell nucleus is reduced when 2 ng (F) or 4 ng (G) of the translational morpholino is injected into zebrafish embryos. (H) lhx5-splicing morpholino reduces wild-type lhx5 transcripts. Arrow indicates wild-type lhx5 or odc1 transcripts. Band above the wild-type lhx5 transcript is caused by splicing with a cryptic GT located in lhx5 intron 3 downstream from the canonical GT site at the exon 3/intron 3 junction; it results in a frame shift. Amplifications of odc1 transcripts serve as controls. EXPRESSION / LABELING:
|
|
Lhx5, Sfrp1a and Sfrp5 regulate the size of the forebrain. Quantitative measurement of forebrain sizes. (A,B) An uninjected control embryo (A) and an lhx5 mRNA-injected embryo (B) are shown to illustrate how quantitative measurements are carried out. Injected embryos and uninjected control embryos were fixed and processed for in situ hybridization with ptc1 probes. Images of labeled embryos were analyzed with Axiovision software (Carl Zeiss). Two parameters were measured for each image. Green double arrow (head thickness) crosses the apex of forebrain and is approximately perpendicular to the tangent plane at the apex. Forebrain length was defined as the distance between the anterior limit and the diencephalic bending point of ptc1 labeling (red stars). The ptc1 bending point is located at the zona limitans intrathalamica that separates the anterior diencephalon and the posterior diencephalon. (C,D) Measurements were normalized by dividing by the values of uninjected controls and multiplying by 100. ** denotes significant differences between uninjected control and affected injected embryos (t-test, P<0.01). For each measurement, the sample size was n=10. EXPRESSION / LABELING:
|
|
sfrp1a promoter fragments drive GFP expression in forebrain regions. Panels show GFP fluorescence from live embryos at the Prim-5 stage (24 hours post-fertilization). |