- Title
-
Blood stem cell-forming haemogenic endothelium in zebrafish derives from arterial endothelium
- Authors
- Bonkhofer, F., Rispoli, R., Pinheiro, P., Krecsmarik, M., Schneider-Swales, J., Tsang, I.H.C., de Bruijn, M., Monteiro, R., Peterkin, T., Patient, R.
- Source
- Full text @ Nat. Commun.
|
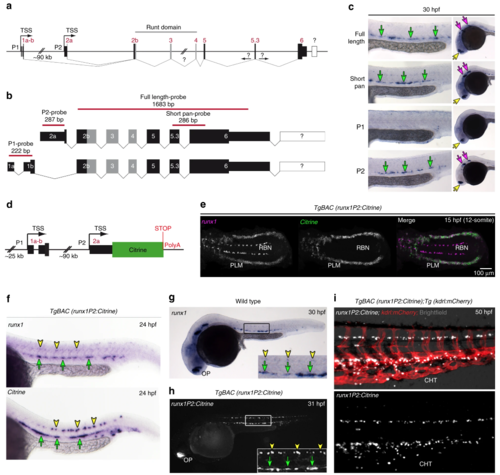
Generation of a zebrafish BAC transgenic reporter line for runx1. a Schematic representation of the zebrafish runx1 locus. Exon nomenclature adjusted to the human RUNX1 locus. P1 and P2 indicate the distal and the proximal promoter respectively (TSS: transcriptional start site). b Schematics of the two alternative transcripts derived from the alternative promoters P1 and P2. Transcript-specific and pan binding ISH probes are indicated with their respective lengths. c ISH for runx1 isoforms in 30 hpf embryos. Left panel: trunk region. Right panel: head region. Green arrows point to the HE, yellow arrows show the olfactory placode and purple arrows depict neurons in the brain region. d Schematic of the recombineered 97a02 BAC. A Citrine reporter cassette was placed downstream of the P2 ATG. e Confocal microscopy image of a flat mounted 15 hpf embryo after double FISH for runx1 and Citrine in TgBAC(runx1P2:Citrine) depicting the region of the posterior lateral plate mesoderm (PLM) and Rohon-Beard neurons (RBN). Maximum intensity projection of a 58 µm stack. f–h Gene expression analysis for runx1and runx1P2:Citrine during definitive haematopoiesis. Green arrows point to the HE. Yellow arrowheads point to neurons in the spinal cord. f ISH for runx1 or Citrine in 24 hpf TgBAC(runx1P2:Citrine) embryos. g ISH for runx1 in 30 hpf embryos. hRepresentative fluorescent microscopy image of a 31 hpf TgBAC(runx1P2:Citrine) embryo. Insets in (g and h) enlarge the boxed region. i Fluorescent microscopy image of a 50 hpf double transgenic TgBAC(runx1P2:Citrine);Tg(kdrl:mCherry) embryo focusing on the region of the caudal haematopoietic tissue (CHT) EXPRESSION / LABELING:
|
|
FACS based isolation of haemogenic and aortic roof endothelium. a Schematic of transgene expression in the trunk region of double transgenic TgBAC(runx1P2:Citrine);Tg(kdrl:mCherry) embryos during definitive haematopoiesis. b Representative confocal image of the DA region of double transgenic TgBAC(runx1P2:Citrine);Tg(kdrl:mCherry) embryos at 29 hpf. Laser intensities enhanced to detect Citrine fluorescence in the DA roof and sprouting inter-somitic vessels (ISV) (orange arrows) in addition to the HE (green arrow). Yellow arrow heads: neurons in the spinal cord. c Characteristic FACS plot of the established gating strategy for the isolation of endothelial sub-populations (runx1hikdrl+ [DP-R1hi]; runx1medkdrl+ [DP-R1med]; runx1lokdrl+[DP-R1lo]; runx1−kdrl+ [SP-kdrl]) from double transgenic TgBAC(runx1P2:Citrine);Tg(kdrl:mCherry) embryos. d qRT-PCR analysis of haematopoietic (tal1, cmyb and gfi1aa) and endothelial (tbx20, efnb2a and dll4) marker gene expression in the different cell fractions isolated from ~29 hpf embryos following the FACS gating strategy depicted in (c). Graphs show the measured mean fold change relative to the expression detected in the DP-R1lo fraction. n = 4 independent experiments. Error bars represent the SEM. 1-way ANOVA; *p < 0.05; **p < 0.01; ***p < 0.001. e Schematic of the vessel structure in the zebrafish trunk region discriminating between aortic roof endothelium (ARE) including the ISV, the HE and the non-aortic endothelium (NAE) including but not restricted to endothelial cells from the cardinal vein (CV). The depicted colour code refers to the colours of the FACS gates presented in (c) EXPRESSION / LABELING:
|
|
Identification of Runx1-regulated genes within the aortic HE. a Experimental scheme to identify Runx1-regulated genes in the HE (DP-R1hi). Highlighted cell populations (DP-R1hi and DN) from un-injected (control) and runx1 MO-injected double transgenic TgBAC(runx1P2:Citrine);Tg(kdrl:mCherry) embryos (28–30 hpf) were used for differential gene expression analysis (DEG) by RNA-seq similar to Fig. 3. b Multidimensional scaling plot (MDS-plot) analysis of the cell populations depicted in (a). cGene list intersection between ARE-specific and Runx1-repressed genes. d Gene list intersection between HE-specific and Runx1-activated genes. e Experimental strategy to analyse potential HE-specific Runx1 target genes by qRT-PCR in runx1+/+(WT) and runx1−/− mutant (MUT) embryos. f Timeline highlighting the time-points (red circles) used for subsequent qRT-PCR analysis. g qRT-PCR gene expression analysis of potential Runx1 targets (dnmt3ba, angpt1, irf1b and pik3cd) in the HE and ARE of runx1+/+ (WT) and the HE of runx1−/− mutant (MUT) embryos at the time-points depicted in (f). Graphs show the mean of detected expression levels relative to the geometric mean (GM) of the two housekeeping genes eef1a1l and rpl13a. n = 5 independent biological experiments for WT embryos and n = 6 independent biological experiments for MUT embryos. Error bars represent the SEM. 2-way ANOVA; *p < 0.05; **p < 0.01; ***p < 0.001. h ISH analysis for the same genes analysed in (g) performed in runx1+/+ and runx1−/− embryos |
|
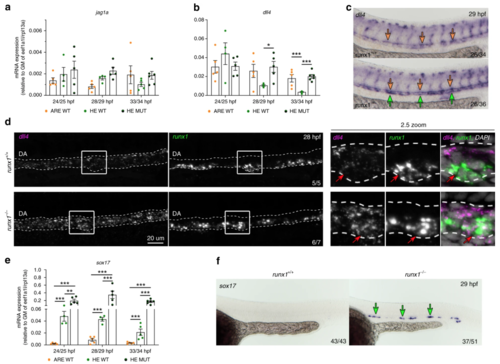
Expression of dll4 in aortic HE is repressed by Runx1. a, b qRT-PCR gene expression analysis of NOTCH ligands jag1a and dll4 in the HE and ARE of runx1+/+ (WT) and the HE of runx1−/− mutant (MUT) embryos. Graphs in (a), (b) and (e) show the mean of detected expression levels relative to the geometric mean (GM) of the two housekeeping genes eef1a1l and rpl13a. n = 5 independent biological experiments for WT embryos and n = 6 independent biological experiments for MUT embryos. Error bars represent the SEM. two-way ANOVA; *p < 0.05; **p < 0.01; ***p < 0.001. c Spatial analysis of gene expression of dll4 in runx1+/+ and runx1−/− embryos by ISH. Green arrows point to the HE. Orange arrows point to the DA roof. d Maximum intensity projection of representative confocal images of runx1+/+ and runx1−/− embryos with double FISH for dll4 and runx1. Red arrows in the close-up highlight the region of a runx1+ cell. e qRT-PCR gene expression analysis of sox17 in the HE and ARE of runx1+/+ (WT) and the HE of runx1−/− mutant (MUT) embryos. f ISH analysis of sox17 in runx1+/+ and runx1−/− embryos. Green arrows point to the HE |
|
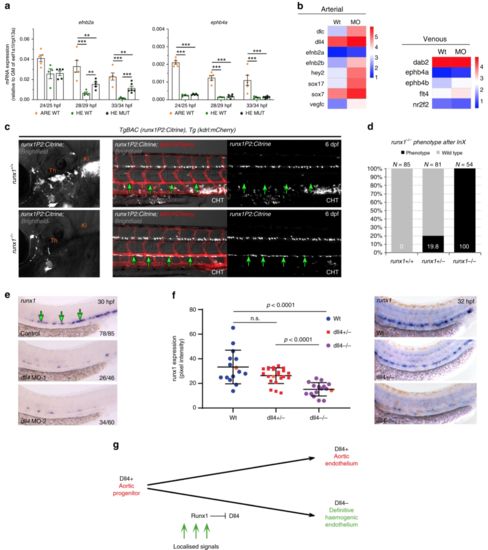
Aortic HE maintains arterial features in the absence of runx1. a qRT-PCR gene expression analysis of the arterial marker efnb2aand the venous marker ephb4b in the HE and ARE of runx1+/+ (WT) and the HE of runx1−/− mutant (MUT) embryos. Graphs show the mean of detected expression levels relative to the geometric mean (GM) of the two housekeeping genes eef1a1l and rpl13a. n = 5 independent biological experiments for WT embryos and n = 6 independent biological experiments for MUT embryos. Error bars represent the SEM. Two-way ANOVA; *p < 0.05; **p < 0.01; ***p < 0.001. b Heatmaps of whole-genome gene expression data from RNA-seq for the HE of control and runx1 MO embryos showing arterial and venous genes. cRepresentative fluorescent microscopy image of 6 dpf TgBAC(runx1P2:Citrine), Tg(kdrl:mCherry) double transgenic embryos on a runx1+/+ or runx1−/− genetic background. Left: Region depicting the region of definitive haematopoietic niches including the thymus (Th) and the kidney (Ki). Right: region depicting the DA and the beginning of the caudal haematopoietic tissue (CHT). Green arrows point to the ventral wall of the DA. d Quantification of embryos derived from the indicated in-crosses (InX) depicting the runx1−/− mutant phenotype as depicted in (c). e ISH analysis of runx1 in control and dll4 MO embryos. Experiment was performed with two different MOs. Green arrows point to the HE. f Offspring of dll4+/− heterozygous fish were analysed by ISH for runx1 at 32 hpf. Runx1 expression was quantified by image analysis and subsequent genotyping57. Representative examples of runx1 expression in each of the WT, dll4+/− and dll4−/− genotypes are shown in the right-hand panels and the corresponding expression values are shown as orange points in the graph. (μwt = 33.3, μmut = 15.19, p < 0.0001, F(2,50) = 16,79, one way ANOVA). g Model depicting the lineage relationship between the definitive HE giving rise to HSCs and arterial endothelium in the DA during embryogenesis. Dll4+ aortic progenitors get exposed to localised signals patterning the DA and inducing haemogenic gene expression. Haemogenic specification includes upregulation of runx1 expression and a subsequent Runx1 induced repression of dll4, thus giving rise to Dll4− mature HE |
|
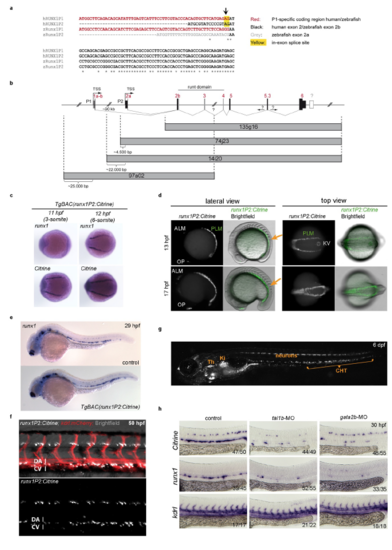
Proper regulation of Citrine expression in TgBAC(runx1P2:Citrine) embryos (a) Multiple sequence alignment between the 5’ most bases of the runx1 alternative promoter products from human (hRUNX1) and zebrafish (zRunx1). P1 specific sequences are shown in red. The zebrafish 2a specific sequence is shown in grey. The in-exon splice site for the human transcript is highlighted in yellow. (b) Schematic representation of the zebrafish runx1 locus and regions covered by each of the previously identified zebrafish BACs1. (c) ISH for runx1 and Citrine in early TgBAC(runx1P2:Citrine) embryos depicting the region of the posterior lateral mesoderm (PLM). (d) Microscopic images of Citrine fluorescence focusing on the PLM region. Orange arrows indicate region shown in the top view panel. (e) ISH for runx1 in WT and early TgBAC(runx1P2:Citrine) embryos at 29 hpf. (f) Fluorescent microscopy image of a 50 hpf double transgenic TgBAC(runx1P2:Citrine);Tg(kdrl:mCherry) embryo focusing on the trunk region. (g) Representative microscopic image of a 6 dpf TgBAC(runx1P2:Citrine) embryo. The region of the thymus (Th), kidney (Ki) and caudal haematopoietic tissue (CHT) are indicated. (h) ISH for Citrine or runx1 in the trunk region of embryos injected with MOs targeting tal1b or gata2b. ISH for kdrl controls for integrity of vascular structures. EXPRESSION / LABELING:
PHENOTYPE:
|
|
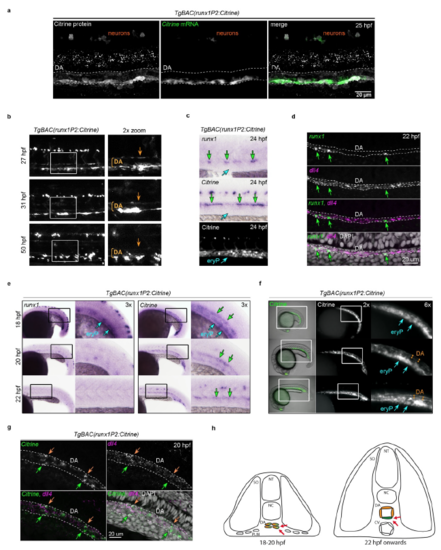
Initial activation of Citrine expression before DA lumenisation causes carry over of Citrine protein in the aortic roof endothelium (a) Confocal image of a whole mount 25 hpf TgBAC(runx1P2:Citrine) embryo immunostained for Citrine protein (white) combined with FISH for Citrine mRNA (green). The DA structure is outlined. (b) Analysis of Citrine transgene and endogenous runx1 expression in 24 hpf TgBAC(runx1P2:Citrine) embryos by ISH (runx1 and Citrine) and fluorescent microscopy (Citrine). Green arrows point to the HE. Blue arrows point to the sub-aortic region containing primitive erythrocytes (eryP). (c) Representative fluorescent microscopic images of the DA region from TgBAC(runx1P2:Citrine) embryos over time. Image acquisition was performed under identical illumination conditions. The magnifications highlight the boxed regions. Orange arrows point to the DA roof. (d) Maximum intensity projection of a representative confocal image after double FISH for dll4 and runx1 in the DA region of a 22 hpf embryo. Green arrows point to runx1+ cells. (e) Expression analysis of runx1 and Citrine during early DA development (18-22 hpf) in TgBAC(runx1P2:Citrine) embryos. Left: ISH analysis of runx1. Right: ISH analysis of Citrine. (f) Fluorescent microscopy images of Citrine fluorescence during early DA development (18-22 hpf) in TgBAC(runx1P2:Citrine) embryos. For e and f: blue arrows point to expression in primitive erythrocytes (eryP); green arrows point to the DA/HE region. Orange double-arrows indicate the DA lumen. (g) Maximum intensity projection of a representative confocal image of double FISH for dll4 and runx1 in the DA region of a 20 hpf embryos. orange arrows point to the ARE and green arrows to the HE. (h) Model depicting the DA region of zebrafish embryos before and after lumenisation. DA angioblasts that reach the embryonic midline are initially clustered together. Subsequent lumenisation creates the geometry for a signalling gradient in which the HE (green) stays in the inductive signalling zone (red arrows) driving the up-regulation of haematopoietic genes. ARE (orange) maintains an arterial programme. CV: cardinal vein; DA: dorsal aorta; NC: notochord; NT: neural tube; SO: somites. |
|
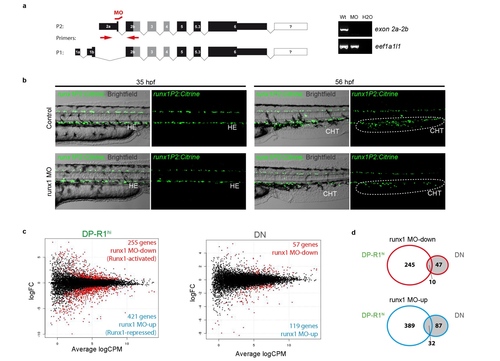
Set-up of a runx1 MO based gene expression screen (a) Left: Schematic of the alternative transcripts derived from the two alternative promoters P1 and P2 and the binding site for the used P2-specific runx1 splice-MO. Right: Typical agarose gel for the analysis of the splice-MO functionality using indicated primers. (b) Fluorescent microscopic images of TgBAC(runx1P2:Citrine) embryos either un-injected (control) or runx1 MO injected. Left: Showing the trunk region including the DA of 35 hpf embryos. Right: Showing the caudal haematopoietic tissue (CHT) region of 56 hpf embryos. (c) Smear plot result of differentially expressed gene analysis (DEG) between runx1 MO and control embryos. Left: DEG analysis for cells of the DP-R1hi gate showing significantly down-regulated (red; Runx1-activated) and up-regulated (blue; Runx1 repressed) genes. Right: DEG analysis for cells of the DN gate showing significantly down-regulated (red) and up-regulated (blue) genes. (d) Gene list intersection between DEG detected in cells of the DP-R1hi and DN gates after runx1 MO injection. Top: Intersection between genes down-regulated due to runx1 MO injection in both populations. Bottom: Intersection between genes down-regulated due to runx1 MO injection in both populations. |
|
Down-regulated expression of dll4 in HE accounts for a lower Dll4 to jag1a ratio driving haemogenic specification (a,b) Spatial analysis of gene expression by ISH. Green arrows point to the HE. Orange arrows point to the DA roof. (a) Time line analysis of dll4 during definitive haematopoiesis in the DA region. The magnifications highlight the boxed regions. (b) ISH analysis of dll4 in runx MO and control embryos. (c) qRT-PCR gene expression analysis of sox32 in the HE and ARE of runx1+/+ (WT) and the HE of runx1-/- mutant (MUT) embryos. Graph shows the mean of detected expression levels relative to the geometric mean (GM) of the 2 housekeeping genes eef1a1l and rpl13a. n = 5 independent biological experiments for WT embryos and n = 6 independent biological experiments for MUT embryos. Error bars represent the SEM. 2-way ANOVA; * = p<0.05; ** = p<0.01; *** = p<0.001. (d) Model depicting the dynamics of NOTCH ligands jag1a |
|
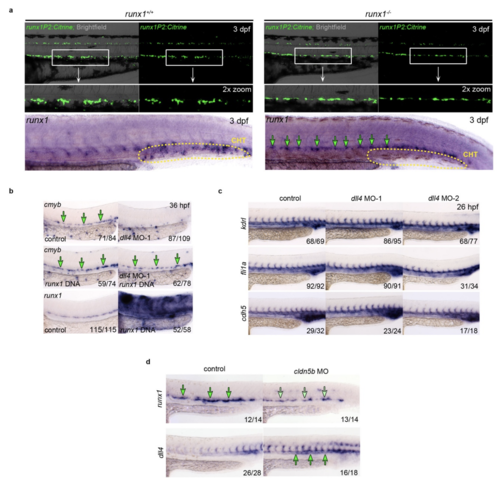
Representative fluorescent microscopy images highlighting the DA region of 3 dpf TgBAC(runx1P2:Citrine) embryos either on a runx1+/+ or runx1-/- genetic background. Magnifications highlight the boxed areas. Bottom: ISH analysis of runx1 in the trunk region of 3 dpf runx1+/+ and runx1-/- embryos. Green arrows point to expression in the HE. The region of the caudal haematopoietic tissue (CHT) is indicated. (b) Top and middle: ISH analysis of cmyb in dll4 MO-1 and control embryos co-injected with or without a runx1 over-expression construct. Green arrows point to the HE. Bottom: Expression of runx1 in control and runx1 over-expression embryos. (c) ISH analysis of endothelial genes kdrl, fli1a and cdh5 in control and dll4 MO embryos. Experiment was performed with two different MOs. (d) ISH analysis of runx1 and dll4 expression in control and cldn5b MO embryos |