- Title
-
Uhrf1 and Dnmt1 are required for development and maintenance of the zebrafish lens
- Authors
- Tittle, R.K., Sze, R., Ng, A., Nuckels, R.J., Swartz, M.E., Anderson, R.M., Bosch, J., Stainier, D.Y., Eberhart, J.K., and Gross, J.M.
- Source
- Full text @ Dev. Biol.
|
uhrf1 mutants possess hypomethylated genomic DNA. Genomic DNA methylation assay in which 750 ng of genomic DNA is digested by a methylation-sensitive enzyme (HpaII), a methylation-insensitive enzyme (MspI), or a mock digestion with no enzyme present. (A) Genomic DNA isolated from Wild-type AB, dnmt1 siblings or dnmt1 mutant embryos demonstrates the efficacy of the assay. Genomic DNA is digested by the methylation-sensitive HpaII to a greater extent in the mutant embryos than in siblings or wild-types. (B) Genomic DNA isolated from Wild-type AB, uhrf1 siblings or uhrf1 mutant embryos; uhrf1 mutant DNA is digested by the methylation sensitive HpaII to a greater extent than sibling or wild-type DNA. (C) SouthWestern assay to quantify 5-methylcytosine levels on genomic DNA. 2 µg of genomic DNA is extracted from the indicated group of embryos, loaded onto a membrane and probed with anti-5-methylcytosine antibody. (D) Quantification of 5-methylcytosine levels (n = 8 trials; *** p < 0.00002). Error bars represent s.e.m. PHENOTYPE:
|
|
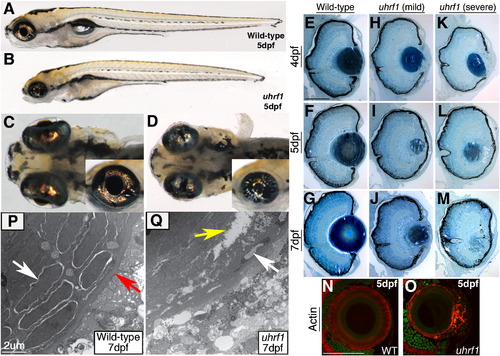
uhrf1 mutants possess abnormal lenses and cataracts. (A,C) Wild-type sibling and (B,D) uhrf1 mutant embryos at 5 dpf. Mutants possess defects in lens formation and cataracts. Pupils are smaller (inset in C,D) and lenses are malformed. (E–M) Transverse histology from wild-type, (H–J) “mild” and (K–M) “severe” uhrf1 mutants at (E,H,K) 4 dpf, (F,I,L) 5 dpf, and (G,J,M) 7 dpf. (H–J) Mild phenotypes include smaller lenses, anterior opacifications and some unraveling of lens fibers at the anterior and/or posterior of the lens. (K–M) Severe phenotypes include substantial opacifications throughout the lens, lens dysplasias, peripheral fiber unraveling from the core of differentiated fibers and lens degeneration. (N,O) Wild-type (N) and mild uhrf1 mutant (O) cryosections stained for F-actin. The uhrf1 anterior lenses contain disorganized, nucleated cells often in excess of the normal wild-type lens epithelial monolayer. (P,Q) TEM analyses of the lens sub-equatorial region in 7 dpf wild-type embryos (P) reveal early differentiating fibers (which still contain nuclei, white arrow) surrounded by the lens capsule (red arrow). In uhrf1 mutants (Q) the lens capsule is absent, and apoptotic lens fiber nuclei (white arrow) and intracellular gaps or tears in the mutant fibers (yellow arrow) are observed. Scale bars are 80 μm. |
|
dnmt1 mutants also possess abnormal lenses and cataracts. (A) Wild-type and (B) dnmt1 mutant embryos at 5 dpf. Mutants display obvious lens defects and in the more severely affected embryos lens dysplasias are observed (arrow in B). F-actin staining of wild-type (A′) and mild dnmt1 (B′) eye cryosections show that the anterior region of dnmt1 mutant lenses contain many disorganized nucleated cells that do not resemble the cuboidal structure of the wild-type lens epithelial monolayer. (C–F) Transverse histology from (C,E) wild-type and (D,F) dnmt1 mutant embryos at (C,D) 5 dpf and (E,F) 7 dpf. Mutants display lens dysplasias, peripheral fiber unraveling from the core of differentiated fibers and lens degeneration. Scale bars are 80 μm. |
|
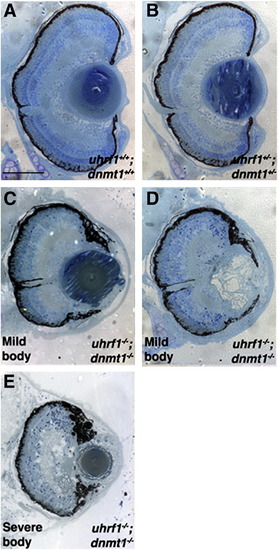
Genetic interaction between uhrf1 and dnmt1 during lens development. (A) 5 dpf histology from uhrf1+/+; dnmt1+/+, (B) uhrf1+/-; dnmt1+/-, and (C–E) uhrf1-/-; dnmt1-/- embryos. No compound heterozygous eye phenotype is observed (B). Lens defects in uhrf1-/-; dnmt1-/- double mutants are no more severe than those in either of the single mutants, and range from mild (C) to severe (D). uhrf1-/-; dnmt1-/- double mutants with severe body morphology (E) do not have more severe lens phenotypes. Scale bars are 80 μm. PHENOTYPE:
|
|
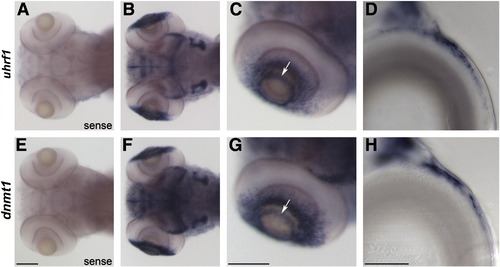
uhrf1 and dnmt1 are expressed in proliferative regions of the lens and retina at the time of phenotype onset. In situ hybridization of uhrf1 sense (A) and antisense (B–D) and dnmt1 sense (E) and antisense (F–H) riboprobe on wild-type embryos at 4 dpf. No expression was detected in sense controls (A,E). Dorsal (B,F) and lateral (C,G) whole mount images demonstrate that uhrf1 and dnmt1 transcripts are expressed in both the ciliary marginal zone of the retina and in the proliferative germinative zone of the lens epithelium (which resembles a ring around the anterior lens when viewed in whole mount, arrows in C and G). Sectioned images of the lens (D, H) confirm the restriction of uhrf1 and dnmt1 expression to distinct regions of the epithelium (anterior is right, dorsal is up). Scale bars in A–C and E–G are 100 μm, and scale bars in D and H are 20 μm. EXPRESSION / LABELING:
|
|
Lens epithelium gene expression, cell proliferation, and apoptosis is altered in uhrf1 and dnmt1 mutant lenses. (A–C,E,F) Whole mount dorsal images of the retina and lens of tgfB3 antisense in situ hybridizations on wild-type (A), uhrf1 (B,E) and dnmt1 (C,F) mutant embryos at 4 dpf. (A) Wild-type lenses showed strong staining for tgfB3, while staining in mutant lenses was classified as either weak (B,C) or moderate (E,F) (summarized in the table (D)). (G–I) Cryosections from 4 dpf embryos immunostained for PCNA (red). PCNA-positive cells are present in the lens epithelium of phenotypically wild-type (G) embryos as well as uhrf1 (H) and dnmt1 (I) mutants. In each panel, arrows of the same color indicate the dorsal and ventral proliferative zones within the lens epithelium, as defined by PCNA staining. (J–L) Cryosections through 4 dpf phenotypically wild-type (J), uhrf1 (K), or dnmt1 (L) lenses stained for incorporation of BrdU (applied between 96 and 98 hpf). White arrows point to lens cells which have incorporated BrdU. Fewer BrdU-incorporating cells are labeled in the uhrf1 and dnmt1 mutant lenses than in wild-type sibling lenses, and mutant cells labeled with BrdU are not positioned exclusively in the lateral epithelium as they are in the wild-type lens. (M–O) Phosphohistone H3 (pH3) immunostaining of 4 dpf (M) phenotypically wild-type, (N) uhrf1 mutant and (O) dnmt1 mutant embryos. White arrows in panel M show two cells in the wild-type lens epithelium stained positively for pH3. (N,O) No pH3 positive cells are observed in mutant lens epithelium. The yellow arrow in (N) indicates a pH3 positive cell in the cornea. Some sections have diffuse background staining in the cornea which is not localized to nuclei. (P–U) TUNEL immunostaining (red) of 4 dpf (P–R) and 5 dpf (S–U) phenotypically wild-type (P,S), uhrf1 (Q,T) and dnmt1 (R,U) mutant cryosections. No apoptotic cells are observed in the lens of wild-type embryos at 4 dpf (P), but numerous TUNEL-positive cells are observed in the still-nucleated epithelial and early fiber regions of the uhrf1 and dnmt1 mutant lenses (white arrows in Q, R). At 5 dpf, TUNEL-positive cells are rare in the wild-type eye (S), however, uhrf1 and dnmt1 mutant lenses contain numerous TUNEL-positive cells in the lens, and some in the cornea and retina (T,U). In all 4 dpf panels, mild mutants were selected for analysis. Sections in panels A–C and J–O are counter-stained with Sytox-Green (green). Anterior is to the right in all panels. Scale bars are 100 μm in A–C and E–F, 70 μm in G–R, and 80 μm in S–O. EXPRESSION / LABELING:
|
|
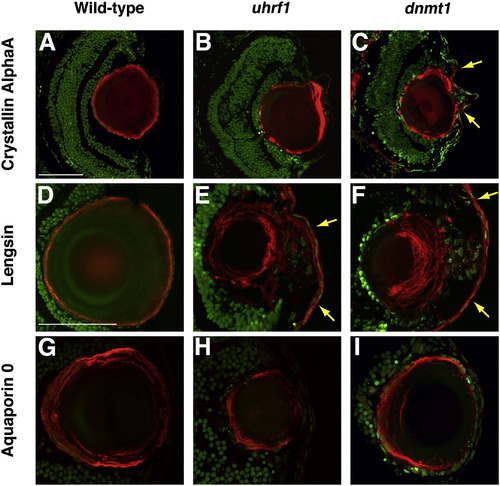
Lens fibers express markers of terminal differentiation in uhrf1 and dnmt1 mutants. (A–I) Cryosections from 5 dpf embryos immunostained for the lens fiber markers Crystallin AlphaA (A–C, red), Lengsin (D–F, red), or Aquaporin 0 (G–I, red). In all sections, nuclei are counterstained with Sytox-Green (green). uhrf1 (B,E,H) and dnmt1 (C,F,I) fibers express these markers, indicating that the disorganized secondary fibers in the mutant lenses still express markers of terminal differentiation. Yellow arrows in panels C, E, and F indicate regions of the mutant cornea which ectopically express lens fiber genes. In all panels, mild mutants were selected for analysis. Anterior is to the right in all panels. Scale bars are 80 μm. EXPRESSION / LABELING:
|
|
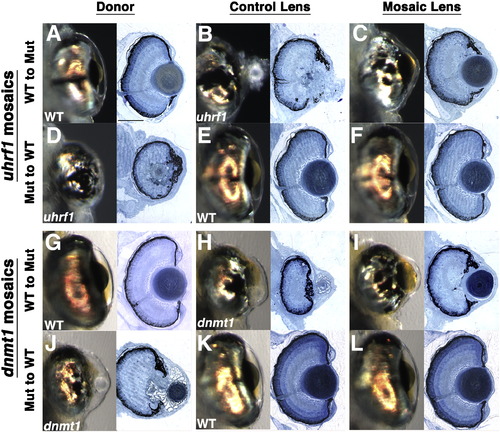
Uhrf1 and Dnmt1 are required lens-autonomously for normal lens development. Shield-stage transplants were performed as described in Fig. S6. (A–F) Representative transplants between wild-type embryos and uhrf1 mutants and (G–L) wild-type embryos and dnmt1 mutants. All images are from 5 dpf embryos and each pair of whole-mount and histology images is derived from the same embryo and eye. For consistency, all data presented are taken from embryos with medium (30–70%) contribution of cells in the mosaic host lens. (A,D,G,J) whole-mount and histology images of eyes formed from donor embryos, (B,E,H,K) whole-mount and histology images of eyes formed from the non-mosaic side of host embryos and (C,F,I,L) whole-mount and histology images of eyes formed from the mosaic lens. Mutant to wild-type (A–C,G–I) transplants and wild-type to mutant transplants (D–F,J–L) both yielded normal lens formation in the mosaic host lens indicating that Uhrf1 and Dnmt1 function is required lens autonomously for lens development. Scale bars are 80 μm. |
|
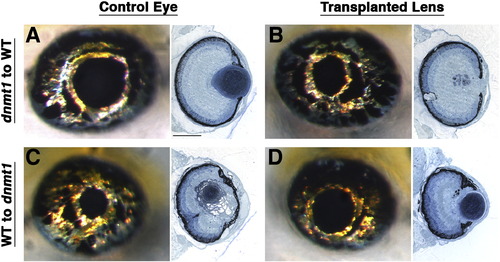
Wild-type retinas cannot rescue development of a mutant lens. Lens transplants performed at 37 hpf (Yamamoto and Jeffery, 2002) demonstrate that mutant lenses placed in wild-type retinas retain the mutant phenotype. (A–D) Representative 5 dpf whole-mount and histology data for lens transplants between wild-type embryos and dnmt1 mutants. Each pair of whole-mount and histology images is derived from the same embryo and eye, and transplants were unilateral so control and transplanted lenses are from the same fish. (A,B) dnmt1 lenses transplanted to wild-type host eyes resulted in the mutant phenotype (n = 5). (C,D) Wild-type lenses transplanted to dnmt1 mutant hosts resulted in wild-type lens formation (n = 5). Similar results were obtained from corresponding transplants between uhrf1 mutant and wild-type embryos (data not shown). Scale bars are 80 μm. |
|
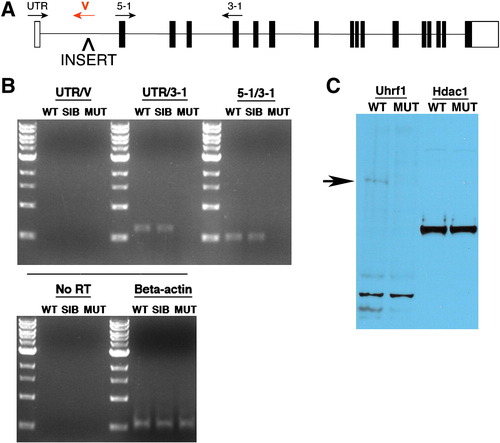
uhrf1hi3020 is a null allele. (A) Schematic of the uhrf1 genomic locus with the approximate position of the proviral insert and viral or exon-specific primers used for RT-PCR analyses indicated. (B) RT-PCR using the indicated primer sets reveals an absence of uhrf1 transcripts in recessive mutants. Beta actin is used as a loading control. (C) Western blot using protein extracts derived from whole, de-yolked 5 dpf wild-type sibling and uhrf1 mutant embryos. Anti-Uhrf1 recognizes a band of approximately 85 kDa in wild-type samples, which is absent in those from uhrf1 mutants. anti-Hdac1 is used as a loading control. EXPRESSION / LABELING:
|
|
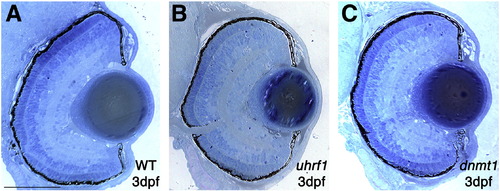
uhrf1 and dnmt1 mutant eyes are phenotypically indistinguishable from wild-type at 3 dpf. Representative histological sections of eyes from wild-type (A), uhrf1 (B) and dnmt1 (C) embryos at 3 dpf. At this point in development, no eye phenotype is observed in the mutant embryos. Scale bars are 60 μm. PHENOTYPE:
|
|
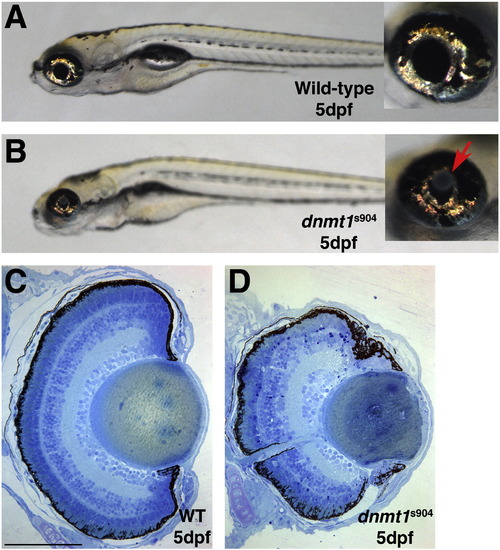
The dnmt1s904 allele phenocopies lens defects in uhrf1 mutants. (A) Wild-type and (B) dnmt1s904 mutant embryos at 5 dpf. As in lenses of uhrf1 and dnmt1s872 mutant embryos, dnmt1s904 mutants display obvious lens defects and in the more severely affected embryos lens dysplasias are observed (arrow in B inset). (C,D) Transverse histology from (C) wild-type and (D) dnmt1s904 mutant embryos at 5 dpf. The mild mutant lens phenotype shown in panel D demonstrates the range of lens defects observed in dnmt1s904 mutants, which is consistent with the range observed in uhrf1 and dnmt1s872 mutants. Scale bars are 80 μm. PHENOTYPE:
|
|
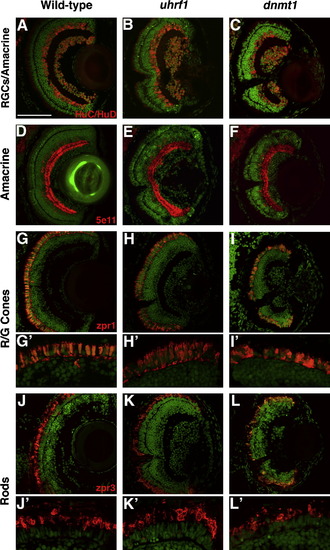
Differentiated retinal cell types are present in uhrf1 and dnmt1 mutants. Cryosections of eyes from wild-type (A,D,G,J), uhrf1 (B,E,H,K) and dnmt1 (C,F,I,L) eyes at 5 dpf. For all panels, antibody staining is in red and sections are counterstained with Sytox-Green (green) to stain nuclei. (A–C) HuC/HuD staining of retinal ganglion cells and amacrine cells. (D–F) 5e11 staining of amacrine cells. (G–I) zpr-1 staining of red/green cones. (J–L) zpr-3 staining of rods. Each of these differentiated cell types is present and in appropriate laminar positions in uhrf1 and dnmt1 mutant retinas. Despite correct localization, both red/green cones and rods have a distorted morphology in uhrf1 and dnmt1 mutant retinas (H′,I′,K′,L′), the severity of which correlated with the severity of lens phenotype. Scale bars are 80 μm. PHENOTYPE:
|
|
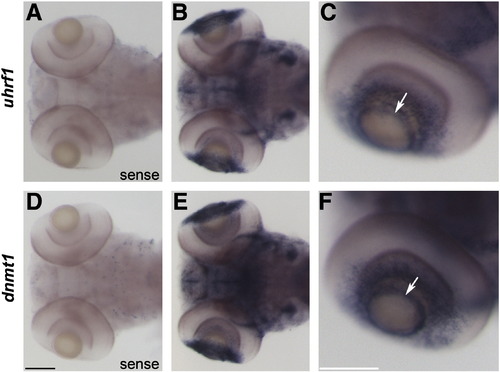
uhrf1 and dnmt1 are expressed in proliferative regions of the lens and retina at 5 dpf. In situ hybridization of uhrf1 sense (A) and antisense (B,C) and dnmt1 sense (D) and antisense (E,F) riboprobe on wild-type embryos at 5 dpf. No expression was detected in sense controls (A,D). Dorsal (B,E) and lateral (C,F) whole mount images demonstrate that uhrf1 and dnmt1 transcripts are present in both the ciliary marginal zone of the retina and in the proliferative germinative zone of the lens epithelium (which resembles a ring around the anterior lens when viewed in whole mount, arrows in C and F). Scale bars are 100 μm. EXPRESSION / LABELING:
|
|
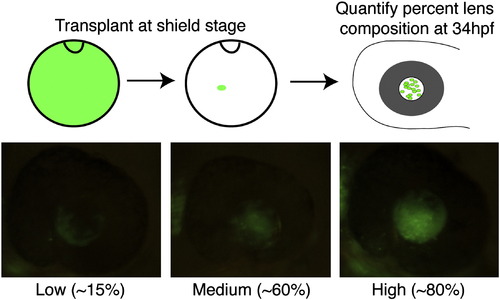
Shield-stage lens mosaics. Cartoon schematic of shield-stage transplants. Donor embryos are injected with Alexa Fluor 488 dextran (10 kDa) and grown to the shield stage. Animal pole view of shield-stage host embryos (shield is located at 12 o′clock in the cartoon), indicating the approximate location of the lens-fated region of the embryo into which donor cells were transplanted. Embryos were grown to 34 hpf and screened for percent lens composition and classed into “Low” (< 30% donor cells), “Medium” (30–70% donor cells) or “High” (> 70% donor cells) groups. Embryos with any retinal contamination were discarded. The genotype of donor and host embryos was determined at 5 dpf based on phenotype, and only those involving severe mutants were used for subsequent analyses. |
|
Control transplants. (A–C) Wild-type to wild-type transplants. (A) Whole mount and histology images of donor eye, (B) host, non-mosaic eye and (C) mosaic lens. All wild-type to wild-type transplants yielded perfectly wild-type lenses. (D–F) uhrf1 to uhrf1 transplants. (D) Whole mount and histology images of donor eye, (E) host, non-mosaic eye and (F) mosaic lens. uhrf1 to uhrf1 transplants yielded mutant lenses. Scale bars are 80 μm. |
|
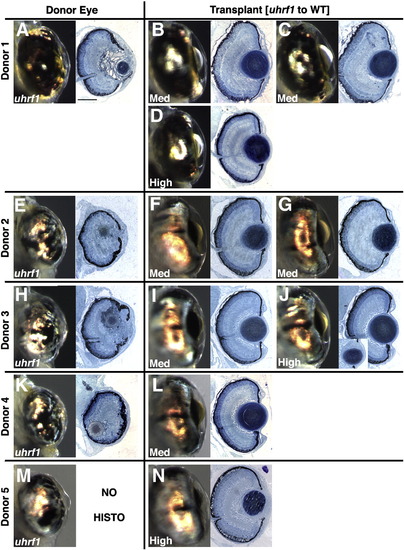
Additional uhrf1 to wild-type transplants. (A,E,H,K,M) Whole mount and histology images of donor eye. (B,C,D,F,G,I,J,L,N) Whole mount and histology images of mosaic lens. Each donor was used to populate one or more hosts as indicated, and the percent mosaic lens composition of donor cells is indicated on each whole-mount image. Mosaic lenses were scored as (B,C,D,F,G,I,L) normal (no phenotype), or (J,N) mild. Inset in (J) is a deeper section through the eye, at the periphery of the lens, where mild defects were noted. Scale bars are 80 μm. |
|
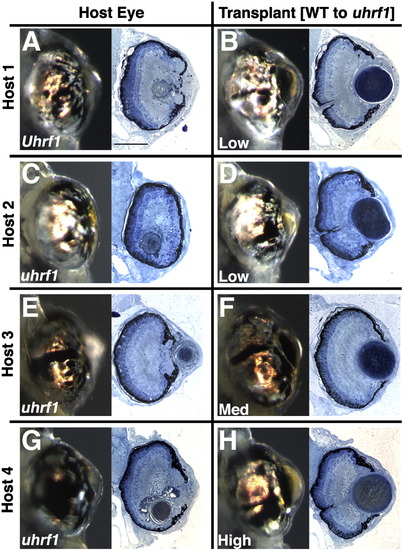
Additional wild-type to uhrf1 transplants. Donors were wild-type siblings. (A,C,E,G) Whole mount and histology images of non-mosaic eye of host embryo. (B,D,F,H) Whole mount and histology images of mosaic lens. The percent mosaic lens composition of donor cells is indicated on each whole-mount image. Mosaic lenses were scored as (D,F,H) normal (no phenotype), or (B) mild. Scale bars are 80 μm. |
|
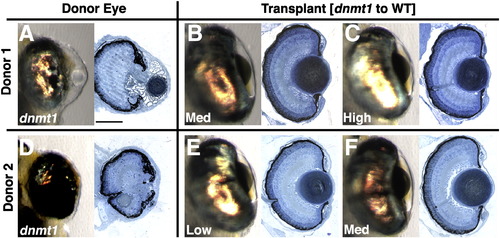
Additional dnmt1 to wild-type transplants. (A,D) Whole mount and histology images of donor eye. (B,C,E,F) Whole mount and histology images of mosaic lens. Each donor was used to populate one or more hosts as indicated, and the percent mosaic lens composition of donor cells is indicated on each whole-mount image. Mosaic lenses were scored as (B,C,E,F) normal (no phenotype). Scale bars are 80 μm. |
|
Additional wild-type to dnmt1 transplants. Donors were wild-type siblings. (A,C,E) Whole mount and histology images of non-mosaic eye of host embryo. (B,D,F) Whole mount and histology images of mosaic lens. The percent mosaic lens composition of donor cells is indicated on each whole-mount image. Mosaic lenses were scored as (B,D,F) mild. Inset in (D) is a deeper section through the eye, at the periphery of the lens, where mild defects were noted. Scale bars are 80 μm. |
|
Wild-type donor cells do not outcompete and repopulate the lens epithelium of a uhrf1 host. Shield-stage transplants were performed in which donor cells were derived from transgenic zebrafish expressing beta actin2:mCherry. uhrf1 mutants or phenotypically wild-type siblings were the transplant hosts. The resulting mosaic embryos were cryosectioned, stained with an anti-dsRed antibody, and imaged by confocal microscopy to visualize beta actin2:mCherry-expressing donor-derived cells within the host lens. (A,B) control (wild-type donor → wild-type host) or (C,D) experimental (wild-type donor → uhrf1 host) mosaic lenses at 4 dpf. In both cases, the lens epithelium contained only a few donor-derived cells. For each mosaic embryo, both the non-transplanted (fully host-derived) lens (A,C), and the mosaic lens (B,D) are shown. Scale bars are 70 μm. |

Unillustrated author statements PHENOTYPE:
|
Reprinted from Developmental Biology, 350(1), Tittle, R.K., Sze, R., Ng, A., Nuckels, R.J., Swartz, M.E., Anderson, R.M., Bosch, J., Stainier, D.Y., Eberhart, J.K., and Gross, J.M., Uhrf1 and Dnmt1 are required for development and maintenance of the zebrafish lens, 50-63, Copyright (2011) with permission from Elsevier. Full text @ Dev. Biol.