Fig. 5
- ID
- ZDB-FIG-260217-16
- Publication
- Chiu et al., 2026 - Glial betaPix is essential for blood vessel development in the zebrafish brain
- Other Figures
- All Figure Page
- Back to All Figure Page
|
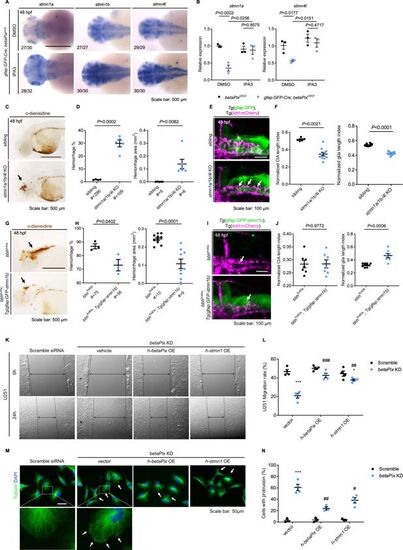
Stathmin acts downstream of betaPix in glial migration via regulating tubulin polymerization. (A) Whole-mount RNA in situ hybridization showing that stmn1a, stmn1b, and stmn4l expression in gfap:GFP-Cre; betaPixct/ct embryos were partially rescued by Pak1 inhibitor IPA3 treatment at 48 hpf. Dorsal views, anterior to the left. (B) qRT-PCR analysis showing that stmn1a and stmn4l expression were rescued in gfap:GFP-Cre; betaPixct/ct mutants by IPA3 treatment at 48 hpf. Each dot represents one embryo. Data are presented in mean ± SEM; one-way ANOVA with Dunnett’s test, individual p-values mentioned in the figure. (C) Representative stereomicroscopy images of erythrocytes stained with o-dianisidine in siblings and stmn1a/1b/4l CRISPR mutants at 48 hpf. Brain hemorrhages, indicated with arrows, appeared in stmn1a/1b/4l mutants. Lateral views with anterior to the left. (D) Quantification of hemorrhagic parameters in (C). Left panel showing hemorrhage percentages, with independent experiment as dot. Right panel showing hemorrhage areas with each dot representing one embryo. # represents the numbers of embryos scored for each analysis, three or more individual experiments conducted. Data are presented in mean ± SEM; unpaired Student’s t-test with individual p-values mentioned in the figure. (E) 3D reconstruction of glial structure (green) and vasculature (magenta) in the heads at 48 hpf. Lateral view with anterior to the left. CtA defects (white arrows) indicated in stmn1a/1b/4l mutants. (F) Quantification of CtA and glia parameters in (E). Length index normalized to individual head length, with each dot representing one embryo. Data are presented in mean ± SEM; unpaired Student’s t-test with individual p-values mentioned in the figure. (G) Representative stereomicroscopy images of erythrocytes stained with o-dianisidine in bbhfn40a and bbhfn40a; Tg(gfap:GFP-stmn1b) embryos at 48 hpf. Brain hemorrhages, indicated with arrows, decreased in bbhfn40a mutants with glia-specific overexpression of stmn1b, compared with bbhfn40a mutant siblings. Lateral views with anterior to the left. (H) Quantification of hemorrhagic parameters of (G). Left panel showing hemorrhage percentages, with independent experiment as dot. Right panel showing hemorrhage areas with each dot representing one embryo. # represents the numbers of embryos scored for each analysis, three or more individual experiments conducted. Data are presented in mean ± SEM; unpaired Student’s t-test with individual p-values mentioned in the figure. (I) 3D reconstruction of the gfap:GFP-stmn1b overexpression (green) and vasculature (magenta) in the heads of bbhfn40a siblings and bbhfn40a; Tg(gfap:GFP-stmn1b) mutants at 48 hpf. Lateral view with anterior to left. White arrows indicate CtA defects. (J) Quantification of CtA and glia parameters in (I). Length index normalized to individual head length, with each dot representing one embryo. Data are presented in mean ± SEM; unpaired Student’s t-test with individual p-values mentioned in the figure. (K) Representative stereomicroscopy images of U251 cells at 0 and 24 hours after wounding. U251 cells were transfected with negative control siRNA or betaPIX siRNA separately, in combination with pcDNA3.1 vector, betaPIX overexpression plasmid, and STMN1 overexpression plasmid. The wound edges are highlighted by dashed lines, with arrow lines indicating the wound width. (L) Quantification of wound closure in (K), showing *p<0.05, ***p<0.005 compared to negative control siRNA with empty vector transfection. ##p<0.01, ###p<0.005 compared to betaPix knockdown with empty vector transfection. Data are presented in mean ± SEM; one-way ANOVA with Dunnett’s test. (M) Representative immunofluorescence image of alpha-tubulin and DAPI signals in U251 cells. U251 cells were transfected with negative control siRNA or betaPIX siRNA separately, in combination with pcDNA3.1 vector, betaPIX overexpression plasmid, and STMN1 overexpression plasmid. Box areas are shown in higher magnifications. Arrows indicate protrusions at the cell periphery. (N) Quantification of cell percentages with protrusions in (M). ***p<0.005 compared to negative control siRNA with empty vector transfection. #p<0.05, ##p<0.01 compared to betaPix knockdown with empty vector transfection. Data are presented in mean ± SEM; one-way ANOVA with Dunnett’s test. Individual scale bars are indicated in the figure. |