- Title
-
Reverse Genetic Screening Reveals Poor Correlation between Morpholino-Induced and Mutant Phenotypes in Zebrafish
- Authors
- Kok, F.O., Shin, M., Ni, C., Gupta, A., Grosse, A.S., van Impel, A., Kirchmaier, B.C., Peterson-Maduro, J., Kourkoulis, G., Male, I., DeSantis, D.F., Sheppard-Tindell, S., Ebarasi, L., Betsholtz, C., Schulte-Merker, S., Wolfe, S.A., Lawson, N.D.
- Source
- Full text @ Dev. Cell
|
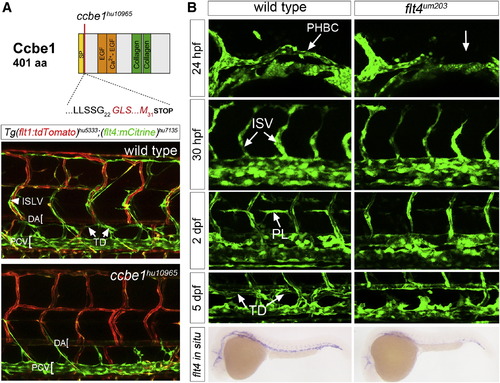
TALEN-Generated Mutations in ccbe1 and flt4 Cause Lymphatic Defects (A) Schematic of Ccbe1 protein indicating position of the ccbe1hu10965 frameshift. SP, signal peptide; EGF, EGF-like domain; Ca2+-EGF, calcium-binding EGF-like domain; and collagen, collagen-like domain. Images are confocal micrographs of wild-type or ccbe1hu10965 mutant embryo bearing Tg(flt4:mCitrine)hu7135 (green) and Tg(flt1:tdTomato)hu5333 (red) transgenes. Homozygous ccbe1hu10965 mutants lacked the TD and ISLVs at 5 days postfertilization; position of the DA and posterior cardinal vein (PCV) are indicated. Lateral views, anterior to the left, and dorsal is up. (B) Confocal micrographs of Tg(fli1a:egfp)y1 wild-type and flt4um203 sibling embryos at 24 hrpf, 30 hrpf, 2 days postfertilization, and 5 days postfertilization. Positions of the PHBC, ISVs, PLs, and TD are indicated. All of these structures except ISVs are missing in flt4um203 mutants at indicated stages (right). (Bottom) flt4 transcript by whole mount in situ hybridization in wild-type and flt4um203 mutant embryos at 28 hrpf. Lateral views, anterior to the left, and dorsal is up. |
|
Normal ISV Development in Selected Mutant Embryos (A?F) Confocal micrographs of Tg(fli1a:egfp)y1 or Tg(kdrl:egfp)s843 transgenic embryos at 32 hrpf subjected to immunostaining with GFP antibody. (A) Wild-type sibling: ISVs, DLAV, and DA are indicated. (B?F) Embryos mutant for (B) arhgef9bum45, (C) fgd5aum134, (D) kalrnbum80, (E) fmnl3um150, or (F) pdgfrbum148. Lateral views, anterior to the left, and dorsal is up. (G) Whole mount in situ hybridization analysis of fmnl3 wild-type and fmnl3um150 mutant sibling at 26 hrpf. Lateral views, anterior to the left, and dorsal is up. (H) Schematic of fmnl3 locus flanking um150 allele and genotyping of individual embryo heads at 72 hrpf from an in-cross of fmnl3um150 heterozygous carriers. Homozygous mutants and heterozygotes pooled for subsequent RT-PCR are indicated by M and het, respectively. (I) Schematic of fmnl3 transcript flanking um150 and RT-PCR analysis of trunks from pooled M and het embryos shown in (H). (J) Whole mount in situ hybridization analysis of pdgfrb in wild-type and pdgfrbum148 mutant at 72 hrpf. Lateral views, anterior to the left, and dorsal is up. (K) Western blot analysis of Pdgfrb and Egfp in lysates from Tg(fli1a:egfp)y1;pdgfrbum148 of indicated genotype at 72 hrpf. (L) Schematic of ets1 exon 3 bearing the um206 lesion and genotyping of individual embryo heads at 3 days postfertilization from an in-cross of ets1um206 heterozygous carriers. (M) Schematic of ets1 transcript and RT-PCR analysis of trunks from pooled M and het embryos shown in (L). EXPRESSION / LABELING:
PHENOTYPE:
|
|
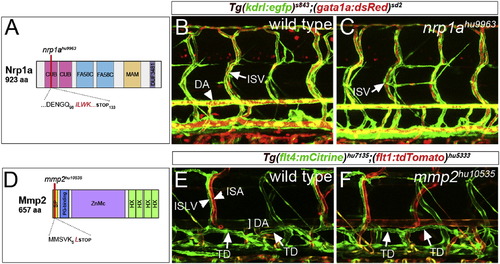
Normal Vascular and Lymphatic Development in nrp1a and mmp2 Mutant Embryos (A) Schematic of Nrp1a domain structure, indicating position of nrp1ahu9963. CUB, CUB domain; FA58C, coagulation factor 5/8 C-terminal domain; MAM, MAM domain; and DUF3481, domain of unknown function. (B and C) Confocal micrograph of embryos bearing Tg(kdrl:egfp)s843 (green) and Tg(gata1a:dsRed)sd2 (red) transgenes at 5 days postfertilization. (B) Wild-type: DA and ISV are indicated. (C) nrp1ahu9963 mutant: ISV carrying red blood cells is indicated. (D) Schematic of the Mmp2 and mmp2hu10535 allele. SP, signal peptide; PG-binding, peptidoglycan-binding domain; ZnMc, zinc-dependent metalloprotease; and HX, hemopexin-like repeats. (E and F) Confocal micrographs of embryos bearing Tg(flt4:mCitrine)hu7135 (green) and Tg(flt1:tdTomato)hu5333 (red) transgenes. (E) Wild-type: ISLV, intersegmental artery (ISA), DA, and TD are indicated. (F) mmp2hu10535 mutant embryo. |
|
Normal Fin Development in fam38a Mutant Embryos (A) Schematic of Fam38a and truncated Fam38aum136. Light blue bars, transmembrane domains; DUF, domain of unknown function. (B and C) Whole mount in situ hybridization of fam38a in (B) wild-type and (C) fam38aum136 mutant at 28 hrpf. (D) Schematic of fam38a exon 8 with um136 lesion and genotyping of individual embryo heads at 72 hrpf from an in-cross of fam38aum136 heterozygous carriers. Homozygous mutants and heterozygotes pooled for subsequent RT-PCR are indicated by M and het, respectively. (E) Schematic of exons in fam38a transcript (numbers in each box indicate exon number, followed by exon size). (F) RT-PCR analysis of fam38a transcript in heterozygous and fam38aum136 mutant trunks pooled from the het and M embryos in (D). (G) Top five sequences are directly from PCR products from genomic DNA of numbered individuals in (D). HpyCH4III site is underlined and the 5-bp um136 deletion is evident. Bottom six sequences are cloned fragments from RT-PCR of pooled fam38aum136 mutant embryos (embryos 7, 13, and 15). (H and I) Transmitted light images of the tail fin fold in live (H) wild-type and (I) fam38aum136 mutant embryos at 72 hrpf. EXPRESSION / LABELING:
PHENOTYPE:
|
|
Normal Hindbrain Development in megamind Mutant Embryos (A) Schematic of the megamind locus in wild-type and megamindum209 mutants. The red and green boxes indicate relative position of the TALEN target sequences, and the MO target sequence is shown. (B) RT-PCR for the megamind and cyrano lincRNAs as well as eef1al1 in wild-type and megamindum209 mutant embryos. RT refers to RNA template without () or with (+) reverse transcription. (C?F) Transmitted light images of the head region in embryos at 48 hrpf. The hindbrain ventricle is indicated with an arrow in each image. Lateral views, anterior to the left, and dorsal is up. (C) Wild-type, uninjected; (D) megamindum209 mutant; (E) wild-type injected with 20 ng megamind conserved site MO; and (F) megamindum209 mutant injected with megamind conserved site MO. |
|
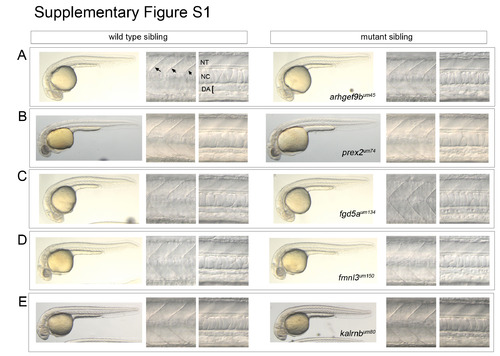
Related to Figure 2. General morphology in selected wild type and mutant sibling embryos. Whole embryos images (left panels in each set of images) were acquired using a standard dissection microscope. Images of somites (middle panels in each set; somite boundaries indicated by arrows in A) and dorsal aorta (DA; right panels in each set) were obtained using differential interference contrast (DIC) microscopy. Position of the neural tube (NT) and notochord (NC) are also noted in (A). All images are lateral views, dorsal is up, anterior to the left. (A) Wild type and arhgef9bum45 mutant sibling embryos. (B) Wild type and prex2um74 mutant sibling embryos. (C) Wild type and fgd5aum134 mutant sibling embryos. (D) Wild type and fmnl3um150 mutant sibling embryos. (E) Wild type and kalrnbum80 mutant sibling embryos. PHENOTYPE:
|

Unillustrated author statements PHENOTYPE:
|
Reprinted from Developmental Cell, 32(1), Kok, F.O., Shin, M., Ni, C., Gupta, A., Grosse, A.S., van Impel, A., Kirchmaier, B.C., Peterson-Maduro, J., Kourkoulis, G., Male, I., DeSantis, D.F., Sheppard-Tindell, S., Ebarasi, L., Betsholtz, C., Schulte-Merker, S., Wolfe, S.A., Lawson, N.D., Reverse Genetic Screening Reveals Poor Correlation between Morpholino-Induced and Mutant Phenotypes in Zebrafish, 97-108, Copyright (2015) with permission from Elsevier. Full text @ Dev. Cell