- Title
-
A spatial and temporal gradient of fgf differentially regulates distinct stages of neural development in the zebrafish inner ear
- Authors
- Vemaraju, S., Kantarci, H., Padanad, M.S., and Riley, B.B.
- Source
- Full text @ PLoS Genet.
|
Development of Statoacoustic Ganglion (SAG). (A) Illustration showing the various stages of SAG development. Neuronal precursors (neuroblasts) are specified (1) and delaminate from (2) the floor of the otic vesicle. Neuroblasts undergo a phase of transit-amplification (3) wherein they migrate to a position between the otic vesicle and hindbrain as they continue to proliferate. Neuroblasts finally differentiate into mature neurons (4). (B) Total number of delaminated neurod+ cells within the SAG counted from serial sections at the indicated times (mean ± standard deviation, n = 2 or greater for each time point). (C) total number of Islet-1-positive SAG neurons at the indicated times (mean of total number ± standard deviation, n = 20 for each time point). (D) neurog1 expression at 30 hpf. (E, F) Co-staining for neurod (blue) and BrdU (red) in embryos exposed to BrdU for 6 hours starting at 26 hpf (E) and 96 hpf (F), and then fixed at 32 hpf and 102 hpf, respectively. (G) Co-staining for isl2b:Gfp (green) and phospho-histone H3 (PH3, red) at 32 hpf. Only one mitotic cell (arrow) is seen in the vicinity of the SAG. (H) Co-staining for Islet1 (green) and BrdU (red) at 36 hpf. Only one double-stained cell is visible (arrow). (I-M) Expression of neurod (blue) and Islet-1 (red) at 36 hpf. Mature neurons are labeled with Islet-1 (I) and delaminated progenitor cells express neurod (J). Positions of section-planes in K–M are indicated in (J). (K-L) Transverse sections passing through the anterior (K), middle (L) and posterior (M) regions of the SAG show mostly complementary patterns of neurod and Islet-1. The outer edge of the otic vesicle is outlined in all panels. SAG cells in stages 1-4 of development are indicated in sections, and the position of the utricular macula (u) is indicated. Images of whole-mount specimens (G, I, J) show dorsolateral (G, I) and dorsal (J) views with anterior to the left. Images of transverse sections (C-F, H, K-M) show dorsal to the top and lateral to the left. Scale bar, 25 μm. |
|
Mature SAG neurons express fgf5. (A-C) Wholemount embryos showing lateral views of fgf5 expression at 24 hpf (A) and 27 hpf (B) and a dorsal view at 30 hpf (C). During these stages, fgf5 expression marks the trigeminal ganglion (tg), anterior lateral line ganglion (all) and SAG, and there is also weak diffuse expression in the developing hindbrain (hb). There is no detectable staining in the otic vesicle (ov). (D-F) Transverse sections (dorsal to the top and lateral to the left) of specimens co-stained for fgf5 (blue) and Islet-1 (red) at 24 hpf (D), 30 hpf (E) and 48 hpf (F). Sections pass through the middle portion of the SAG at the level of the utricular macula (u). The inner and outer surfaces of the otic vesicle are outlined. Co-labeling confirms that fgf5 expression in the SAG is restricted to mature neurons. Scale bar, 25 μm. During mid-somitogenesis stages fgf5 is diffusely expressed throughout the neural tube and strongly marks the developing trigeminal ganglion (not shown). |
|
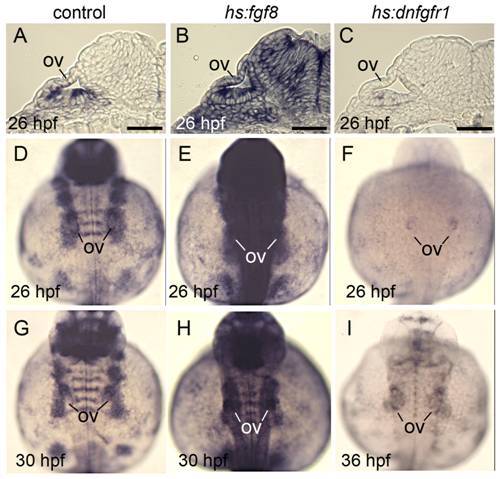
Effects of transgene activation on expression of the Fgf reporter etv5b. All embryos were heat shocked for 30 minutes beginning at 24 hpf. Wild-type and hs:fgf8 embryos were heat shocked at 39°C and hs:dnfgfr1 embryos were heat shocked at 38°C. (A-C) Cross sections showing etv5b expression in wild-type (A), hs:fgf (B) and hs:dnfgfr1 (C) embryos at 26 hpf. (D-I) Dorsal views (anterior up) of wholemounts showing etv5b expression in wild-type (D, G), hs:fgf8 (E, H) and hs:dnfgfr1 (F, I) embryos at the indicated times. The otic vesicle (ov) is marked. Expression of etv5b remains elevated in hs:fgf8 embryos for at least 6 hours after heat shock, whereas etv5b expression is downregulated in hs:dnfgfr1 for at least 12 hours after heat shock. Scale bar, 25 μm. |
|
Fgf regulates neuroblast specification. The icon at the top of the figure indicates that analysis focuses on initial stages of neuroblast formation, as normally marked by neurog1 expression. Experimental manipulations in groups I, II and III are briefly summarized at the tops of the corresponding data panels. (A-C) Experiment I, neurog1 expression at 18 hpf in a control (A), SU5402 inhibitor treated (B) and hs:dnfgfr1/+ transgenic embryo heat shocked for 30 minutes at 38°C beginning at 14 hpf. Blocking Fgf strongly reduces expression of neurog1. (D-G) Experiment II, expression of neurog1 (D-E) and hmx3 (F-G) in control and hs:fgf8/+ embryos heat shocked at 35°C for 6 hours, from 18 hpf until 24 hpf. This regimen results in weak overexpression of Fgf8, which at this stage enhances expression of neurog1. (H) Experiment II, total number of Islet1-positive cells in the SAG (mean and standard deviation, n = 15) at 42 hpf following heat shock activation of hs:fgf8 at 18 hpf at indicated temperatures. Weak misexpression of Fgf8 (35°C) increases production of SAG neurons whereas strong misexpression of Fgf8 (39°C) reduces production of SAG neurons. *p<0.001 in comparison to the control, analyzed with Student′s t test. (I–K) Group III, neurog1 expression at 30 hpf following heat shock at 24 hpf in control embryos (I), hs:fgf8/+ embryos heat shocked at 39°C for 30 minutes to strongly over-express Fgf (J) and hs:dnfgfr1/+ embryos heat shocked for 2 hours at 35°C and then shifted to 33°C to maintain low level inhibition of Fgf signaling (K). At this stage, weak impairment of Fgf enhances neurog1 expression, consistent with the idea that Fgf levels normally increase during development and become inhibitory for neuroblast specification. All images show dorsolateral views with anterior to the left, and the otic vesicle is outlined. EXPRESSION / LABELING:
|
|
Efficacy of fgf5 splice-blocking MO. (A) Schematic of fgf5 mRNA showing intron-exon structure (not to scale). Binding sites for splice-blocking morpholino at intron1-exon2 junction (I1E2-MO) and PCR primers for RT-PCR (forward P1, reverse P2) are shown. (B) RT-PCR results showing the efficacy of I1E2-MO. fgf5 transcript levels are severely reduced in fgf5 morphants at 24 hpf. odc transcript level was used as a constitutive control. |
|
fgf5 from mature neurons terminates the phase of neuroblast specification. The icon at the top of the figure indicates that analysis focuses on initial formation of neuroblasts. Experimental manipulations in groups IV and V are briefly summarized at the tops of the corresponding data panels. (A-H) Expression of neurog1 in control embryos (A-C), a hs:fgf8 embryo (D), fgf5 morphants (E-G), and a hs:fgf8 embryo injected with fgf5-MO (H) at the indicated stages. Transgenic embryos (D, H) were heat shocked for 30 minutes at 39°C beginning at 24 hpf. Vertical lines in (A, C-E, G, H) indicate the plane of transverse sections in (B, F, and insets in C, D, G and H). (I-L) Expression of isl2b:Gfp at 22 hpf (I, J) and neurog1 at 30 hpf (K, L) in a specimen in which mature (fgf5-expressing) neurons were laser-ablated. The same specimen is shown in all panels. Mature SAG neurons expressing isl2b:Gfp were serially ablated on the left side at 22 hpf (J) and 25 hpf (not shown), and the embryo was fixed and sectioned at 30 hpf to examine neurog1 expression (L). Images of the unablated right side (I, K) were inverted to facilitate comparison. The surface of the otic vesicle is outlined in all panels. Arrows in sections indicate the edges of neurog1 domain in the otic floor. Note that the amount and duration of delamination of neurog1+ neuroblasts is strongly enhanced by knockdown of fgf5 (F, G) or ablation of mature neurons (L). Activation of hs:fgf8 reverses the effects of fgf5-MO (H). Scale bar, 25 μm. Transverse sections are shown with lateral to the left and dorsal up. Wholemount images show dorsolateral views with anterior to the left. EXPRESSION / LABELING:
|
|
Normal axial patterning in fgf5 morphants. (A-H) Expression of regional patterning markers in control embryos (A–D) and fgf5 morphants (E-H). Expression of pax5 (A, E) and pou3f3b (B, F) labels anterior and posterior regions, respectively. Expression of otx1a (C, G) and dlx3b (D, H) labels ventromedial and dorsolateral regions, respectively. The otic vesicle is outlined. Images show dorsolateral views with anterior to the left. (I) The total number of hair cells in the utricular (ut) and saccular (sac) maculae of control embryos and fgf5 morphants at 32 hpf. Data were obtained by counting GFP-positive hair cells (mean of total number ± standard deviation) in the sensory epithelia of brn3c:Gfp transgenic embryos. Data show means and standard deviations from 20 specimens each. Differences between control and experimental specimens were not statistically significant (p = 0.16 for the utricle, p = 0.67 for the saccule) based on Student′s t tests. EXPRESSION / LABELING:
|
|
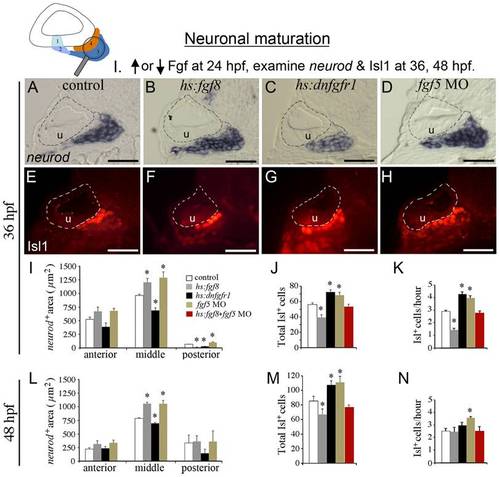
Fgf regulates the balance between transit-amplification and differentiation. The icon at the top of the figure indicates that neuronal maturation (neurod+ transit-amplifying cells and Isl1+ mature neurons) is the focus of analysis. Manipulations in these experiments (Neuronal maturation group I) are briefly summarized at the top. Embryos were heat shocked for 30 minutes at 39°C (wild-type controls, hs:fgf8/+ embryos, and fgf5-morphants) or 38°C (hs:dnfgfr1/+ embryos) beginning at 24 hpf. (A–H) Transverse sections (lateral to the left, dorsal up) showing neurod expression (A-D) or Isl1 staining (E-H) at 36 hpf in control embryo (A, E), hs:fgf8/+ embryos (B, F), hs:dnfgfr1/+ embryos (C, G) and fgf5 morphants (D, H). All sections shown pass through the middle region of the SAG at the level of the utricular macula (u). The otic vesicle is outlined. Scale bar, 25 μm. (I-N) Quantitation of transit-amplifying and mature neuronal populations at 36 hpf (I-K) and at 48 hpf (L–N). Panel I shows a color key to facilitate comparison between treatments: White bars, control; gray bars, hs:fgf8; black bars, hs:dnfgfr1; brown bars, fgf5 morphants; red bars, activation of hs:fgf8 in fgf5 morphants. Analysis of transverse sections was used to measure the mean area of neurod+ precursor cells (I, L) in the anterior, middle and posterior regions of SAG. The total number of Isl1+ neurons (J, M) and the mean hourly rate of neuron production from 24 hpf to 36 hpf (K) and from 36 to 48 hpf (N) was measured by counting neurons in stained wholemount specimens. Error bars in I, J, L, M indicate standard deviations (n = 3 or greater for sectional areas; n = 15 for Isl1+ cell counts). *p<0.05 in comparison to control, analyzed with Student′s t test. Error bars in K, N indicate standard errors. EXPRESSION / LABELING:
|