- Title
-
Maternal Interferon Regulatory Factor 6 is required for the differentiation of primary superficial epithelia in Danio and Xenopus embryos
- Authors
- Sabel, J.L., d'Alençon, C., O'Brien, E.K., Otterloo, E.V., Lutz, K., Cuykendall, T.N., Schutte, B.C., Houston, D.W., and Cornell, R.A.
- Source
- Full text @ Dev. Biol.
|
Danio rerio irf6 expression by in situ hybridization. Lateral views, except as indicated, of D. rerio embryos fixed at the indicated stage and processed with an antisense irf6 probe, except B and inset in C. (A) A cleavage stage embryo, revealing high-level maternal expression. (B) A cleavage stage embryo, processed with an irf6 sense probe, revealing very little background staining. (C) Section at shield stage showing low-level expression in deep cells (DEL) and higher level expression in EVL and dorsal frontrunner cells (DFC). Inset, section of a shield stage embryo processed with a sense probe. (D) Diffuse expression throughout the embryo, with increased expression in the nasal and otic placodes (arrowheads). Head is to the left in panels D–F. (E) Decreasing diffuse expression, expression present in nasal and otic placodes and lateral line primordium (arrowhead). (F) Expression as in E but now including putative gut precursor cells (arrowhead). (G) Ventral view, showing expression in pharyngeal pouches (arrowhead). Expression is not detectable in skin at this stage (asterisk) (e, eye). H) Expression in pharyngeal pouches (arrowhead) and ear (asterisk). (I) Horizontal section, showing expression in oral epithelium (arrowhead). (J) Ventral view, expression in pharyngeal pouches (arrowheads). (e, eye). EXPRESSION / LABELING:
|
|
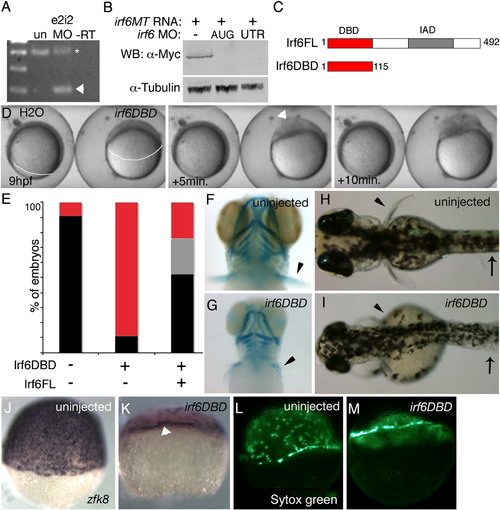
In D. rerio, expression of a putative dominant negative Irf6 causes severe gastrulation defects and late defects in the pectoral fin and skin. (A) RT-PCR analysis of efficacy of splice blocker MO targeting D. rerio irf6. RNA was harvested from 22 hpf embryos that were either uninjected or injected at the single cell stage with 5 ng of irf6 exon 2/intron 2 MO. First strand cDNA was synthesized and PCR carried out with forward primer in exon 1 and reverse primer in exon 3, yielding, from uninjected embryos (un), a single band of expected size, and from MO-injected embryo a second, smaller band (arrowhead). The MO-dependent PCR product was excised and sequenced and found to reflect loss of the first coding exon, which encodes the first 60 amino acids, including half of the predicted DNA binding domain. In e2i2 MO injected embryos, a substantial fraction of irf6 mRNA, possibly corresponding to maternal stores, is spliced correctly (asterisk). - RT, negative control lacking reverse transcriptase (B) Western blot analysis of efficacy of MO targeting translation of Irf6. All lanes show lysates from 16 hpf embryos injected at the two-cell stage with in vitro transcribed irf6 RNA altered to include most of the 5′ UTR and to encode full length Irf6 with a carboxy-terminal Myc tag. -, embryos not injected with MO. AUG, RNA-injected embryos subsequently with 5 ng of irf6 AUG MO; 5′UTR, RNA-injected embryos subsequently with 5 ng of irf6 5′UTRb MO. In a similar Western blot experiment, embryos injected with 20 ng of a 5-base pair mismatch variant of the AUG MO were comparable to embryos not injected with MO (not shown). (C) Schematics of full-length Irf6 and the DNA binding domain variant, which contains the amino terminal 115 amino acids. IAD, Interferon Association Domain (Taniguchi et al., 2001). (D) Frames from a time-lapse video of a water-injected control, left, and an irf6DBD-injected embryo, right, taken at the indicated times. In the latter, thinning of the DEL is delayed (most obvious at the animal pole), as is vegetal migration of the DEL (white line, extent of DEL migration). Finally, when siblings are at about 80% epiboly (9 h), the irf6DBD-injected embryo ruptures near the animal pole. Arrowhead indicates first area of embryo rupture. Staging was analyzed in 10 embryos in three separate experiments with equivalent results. (E) Histogram showing percentage of embryos at 24 hpf of the indicated class that were normal (black), or dead (red), or with head and tail defects (grey). (F, G) Ventral views of 4 dpf embryos stained with alcian-green. (F) Uninjected embryo. (G) irf6DBD-injected embryos showing pharyngeal cartilage elements are present but smaller than normal and pectoral fins (arrowheads) are reduced. (H, I) Dorsal views of live, (H) uninjected embryo and, (I) irf6DBD-injected embryos at 4 dpf. The later shows blistered skin (arrows) and abnormal fins (arrowheads). (J–M) Lateral views of, (J, L) uninjected and, (K, M) irf6DBD-injected embryos at shield stage. (J, K) Embryos processed to reveal expression of zfk8, a marker of the EVL. In the irf6DBD-injected embryo, the vegetal limit of the EVL (arrowhead) is at the position appropriate for dome stage. (L, M) YSL nuclei labeled with sytox green, revealing that in, M) the irf6DBD-injected embryo vegetal migration of the YSL has stalled at dome stage. EXPRESSION / LABELING:
PHENOTYPE:
|
|
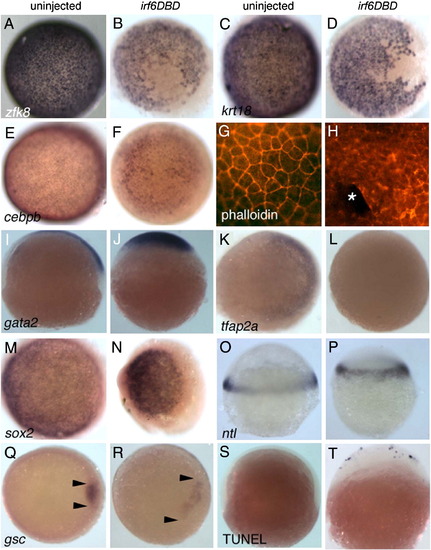
Irf6DBD-mediated changes in gene expression in shield-stage D. rerio embryos. Dorsal views, except as indicated, of uninjected and irf6DBD-injected embryos, as indicated by the column header, fixed when uninjected sibling embryos reached shield stage. Embryos processed for in situ hybridization to reveal expression of the indicated gene, except as noted. Images portray the phenotype displayed by 60%–80% of embryos in the experimental group, n = 25 embryos or greater. (A–F) Expression of zfk8, krt18, and cebpb, are expressed in all EVL cells of uninjected embryos but absent in large patches in uninjected embryos. (G) Uninjected and, (H) irf6DBD-injected embryos stained with phalloidin, revealing polymerized actin (i.e., F-actin). (G) In the uninjected embryo, F-actin is visible in a cortical ring within every EVL cell. (H) In the irf6DBD-injected embryo, there is loss (*) or disorganization of F-actin in superficial cells. (I, J) Lateral views, showing gata2 expression in non-neural epiblast is present but displaced the irf6DBD-injected embryo. (K, L) In contrast to gata2, expression of tfap2a, a marker of non-neural epiblast, is highly reduced in the irf6DBD-injected embryo. (M,N) sox2 expression, a marker of neural epiblast, is present in the irf6DBD-injected embryo. (O, P) Lateral views, showing that ntl expression, a marker of hypoblast, is present in the irf6DBD-injected embryo. (Q, R) gsc expression is present in the shield hypoblast of the uninjected embryo, and in an expanded arc in the irf6DBD-injected embryo, consistent with developmental delay in the latter (arrowheads show extent of gsc expression arc). (S, T) Lateral views of embryos processed to reveal cells undergoing apoptosis by TUNEL. TUNEL-positive cells are absent from this uninjected embryo. In the irf6DBD-injected embryo, many TUNEL-positive cells are visible, primarily within the EVL. |
|
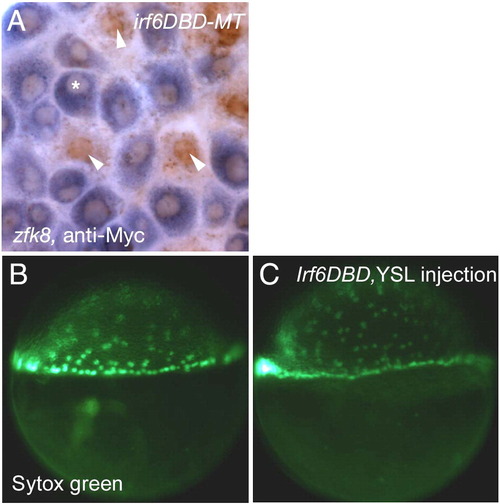
The Irf6DBD effect is cell autonomous to the EVL. (A) Animal pole view of an embryo injected with irf6DBD-MT mRNA into a single blastocyst at 16–32 cell stage, processed at shield stage for zfk8 mRNA (purple) and anti-cMyc IR (brown). Superficial cells which contain anti-Myc IR (arrowheads) have highly reduced zfk8 expression in comparison to cells that do not contain anti-Myc IR (*). (6 cells clearly containing nuclear anti-Myc IR and also clearly in the same plane at the EVL observed in 3 embryos, 0/6 displayed detectable zfk8 signal). (B, C) Lateral views of embryos at shield stage which had been injected with (B) SYTOX green or (C) SYTOX green and irf6DBD into the yolk at sphere stage (4 hpf). There is normal progression of epiboly of yolk syncytial nuclei in both embryos. EXPRESSION / LABELING:
|
Reprinted from Developmental Biology, 325(1), Sabel, J.L., d'Alençon, C., O'Brien, E.K., Otterloo, E.V., Lutz, K., Cuykendall, T.N., Schutte, B.C., Houston, D.W., and Cornell, R.A., Maternal Interferon Regulatory Factor 6 is required for the differentiation of primary superficial epithelia in Danio and Xenopus embryos, 249-262, Copyright (2009) with permission from Elsevier. Full text @ Dev. Biol.