Fig. S5
|
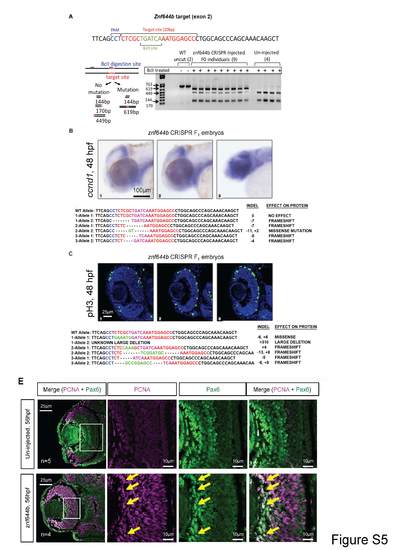
Top: CRISPR-Cas9 targeting of the znf644b gene show multiple mutated alleles in the F1 generation that correlates with cellular defects in the retina. (A) PCR and restriction digest diagnostic on genomic DNA extracted from individual F0 embryos injected with Cas9 mRNA and sgRNA targeting against exon 2 of znf644b. The PCR fragment contains two Bc1I digestion sites, one of which is located within the sgRNAtargeting site. Digestion of wild-type sequences results in three fragments, 449bp, 170bp and 144bp, whereas CRISPR-mediated insertion/deletion mutations result in loss of the digestion site in the target site, resulting in an additional 619bp fragment. (B, top) WISH assays monitoring the expression of ccnd1 in F1 CRISPR embryos at 48hpf. One F1 embryo heterozygous for a frameshift mutation and was phenotypically normal for brain and retinal growth as well as ccnd1 expression at 48 hpf (embryo#1). A separate embryo with two mutant (frameshift) alleles showed significantly reduced brain and retinal growth and elevated expression of ccnd1 (embryo#3), which was reminiscent of the znf644b morphant phenotype. (C, top) Immunostaining assay monitoring pH3-positive cells in retinal cross-sections from F1 CRISPR embryos at 48hpf. One embryo had two znf644b mutant alleles, one of which was predicted to be a very large deletion likely resulting in a non-functional protein, while the other allele had a missense mutation leading to a predicted substitution of only two amino acids. The overall phenotype of this embryo was comparable to WT, suggesting that this embryo was functionally heterozygous (embryo#1). Other embryos genotyped as heteroallelic mutants displayed phenotypes that were similar to the morphants, such as a reduced retinal size and persistent pH3+ cells in the central retina at 48 hpf (embryo#2: 11.67 ± 4.16; embryo #3: 12.67 ± 2.08 compared to WT: 3.8 ± 0.5, n=9). (B & C, bottom) The corresponding sequencing data for both alleles of the embryos analyzed, with the predicted effect on the protein function. Black letters represent unchanged bases; Blue letters represent the PAM sequence; Red letters represent the target sequence; Pink letters represent the Bc1I digestion site; Dashes represent deletions; Green letters represent inserted bases. Bottom: Expression overlap of cell cycle regulator PCNA and the retinal differentiation marker Pax6 in the znf644b morphant retina. At 56 hpf, which coincides with the onset of widespread cell death of central retinal cells in znf644b morphants, PCNA can be found to be expressed in differentiated amacrine and ganglion cells (Pax6-postive) in the znf644b morphant retina, even though this population of neurons is reduced overall. |