- Title
-
Zebrafish rest regulates developmental gene expression but not neurogenesis
- Authors
- Kok, F.O., Taibi, A., Wanner, S.J., Xie, X., Moravec, C.E., Love, C.E., Prince, V.E., Mumm, J.S., and Sirotkin, H.I.
- Source
- Full text @ Development
|
Zinc-finger nuclease-mediated targeting of zebrafish rest. (A) Sequence chromatograms from wild-type siblings (left) and rest mutants (right). The 7 bp deletion (?7) in SBU29 and 4 bp insertion (+4) in SBU34 are illustrated beneath. (B) Domain structure of wild-type Rest and truncated proteins produced by SBU29 and SBU34 mutations. (C) Animal pole views of 6-hpf wild-type and MZrestsbu29/sbu29 embryos. Rest immunoreactivity, TOPRO-3 staining of nuclei, and merged channels are shown. Abundant cytoplasmic Rest protein is detected in wild-type embryos, but not in MZrestsbu29/sbu29 mutants. Confocal images are single 1-μm stacks taken at 40� magnification. EXPRESSION / LABELING:
|

ZFIN is incorporating published figure images and captions as part of an ongoing project. Figures from some publications have not yet been curated, or are not available for display because of copyright restrictions. EXPRESSION / LABELING:
|

ZFIN is incorporating published figure images and captions as part of an ongoing project. Figures from some publications have not yet been curated, or are not available for display because of copyright restrictions. EXPRESSION / LABELING:
|

ZFIN is incorporating published figure images and captions as part of an ongoing project. Figures from some publications have not yet been curated, or are not available for display because of copyright restrictions. EXPRESSION / LABELING:
|
|
Neurogenesis progresses normally in rest mutants. Whole-mount in situ hybridization for the pan-neural marker huC in stage-matched (A,B) 2-somite, (C,D) 8-somite, (E,F) 14-somite and (G,H) 24-hpf wild type or rest heterozygotes and MZrestsbu29/sbu29 mutants. All views are dorsal, anterior to the left. MZrest mutants did not show ectopic or precocious huC expression. To control for subtle differences in developmental timing and genetic background, comparisons were undertaken in crosses of restsbu29/sbu29 females to restsbu29/+ males and between wild-type intercrosses and MZrestsbu29/sbu29 intercrosses. Genotypes were determined by PCR. EXPRESSION / LABELING:
PHENOTYPE:
|
|
Rest function is crucial for the regulation of oligodendrocyte precursor cells in the dorsal spinal cord and for facial branchiomotor migration. (A-D) Lateral views of live 50-hpf and 98-hpf Tg(olig2:GFP) (A,C) and Tg(olig2:GFP); MZrestsbu29/sbu29 mutants (B,D). The number of migrating olig2+ oligodendrocyte precursors (arrows) is significantly reduced in rest mutants at 50 hpf (B) compared with controls (A). The number of migrating OPCs (arrows) in 98-hpf MZrestsbu29/sbu29 mutants (D) is similar to controls (C). (E-G) Dorsal views of the hindbrain of 48-hpf Tg(islet1:GFP) transgenic zebrafish embryos. Representative embryos from a restsbu29/+, restsbu29/+; islet1:GPF intercross. (E) rest+/+ embryo showing normal FBMN migration into r6-r7. (F) FBMNs in restsbu29/+ embryos have only subtle migration defects (arrows). (G) In restsbu29/sbu29 embryos, a significant proportion of FBMNs remain in r4-r5 (bracket). |
|
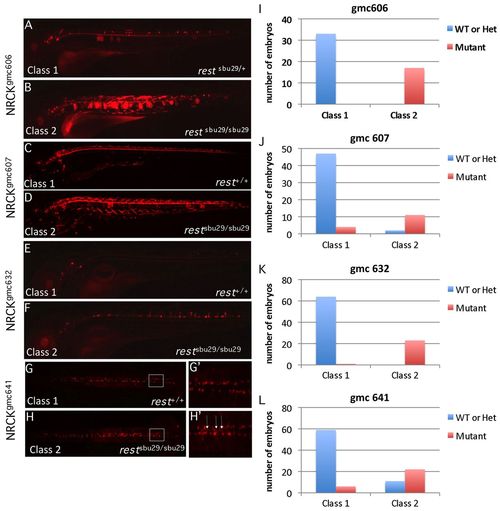
RE1/NRSE reporter transgenic lines are ectopically expressed in rest mutants. (A-H′) Lateral (A-F) or dorsal (G,H) views of live RE1/NRSE transgenic reporter lines in wild-type (A,C,E,G) or restsbu29/sbu29 (B,D,F,H) backgrounds. Expression of each reporter was altered in the mutants. The boxed region in G and H is shown at higher magnification in G′ and H′. Arrows mark midline cells, which are not as numerous in the wild-type background. (I-L) The proportion of embryos in each phenotypic and genotypic class for each reporter. gmc606 and gmc641 larvae are 3 dpf; gmc607, 6 dpf; gmc632, 4 dpf. EXPRESSION / LABELING:
|

ZFIN is incorporating published figure images and captions as part of an ongoing project. Figures from some publications have not yet been curated, or are not available for display because of copyright restrictions. EXPRESSION / LABELING:
|
|
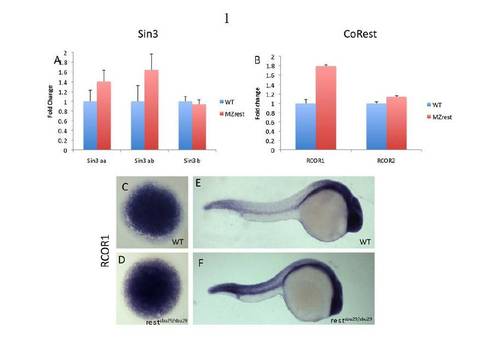
Expression analysis of Rest co-factors in MZrest mutants. (A,B) Expression levels of Rest co-factors assessed by qPCR at mid-blastula stage (4 hpf). Fold differences are relative to wild-type transcript levels (defined as 1). Expression levels of the three zebrafish sin3 homologs and rcor2 are comparable in wild-type and rest mutant embryos. rcor1 mRNA levels are enhanced by 0.75-fold in the rest mutant embryos. Error bars indicate s.e. from three pools of five embryos. (C-F) Whole-mount in situ hybridization (WISH) using rcor1 probe on wild-type and MZrest mutant embryos at 4 hpf (C,D) and 24 hpf (E,F). |
|
Germ layer formation is normal in MZrestsbu29/sbu29 mutants. WISH of germ layer markers in wild-type and rest mutants. All views are dorsal, anterior to the top. (A-F) Domains of mesoderm markers ntl and myod are unaltered in rest mutants. (I,J) Similarly, the endoderm marker axial is also unaltered in rest mutants. (G,H,K,L) Moreover, organizer marker chd and early dorsal neural plate/neural crest marker pax3 expression domains in rest mutants are similar to those of stage-matched wild-type embryos. PHENOTYPE:
|
|
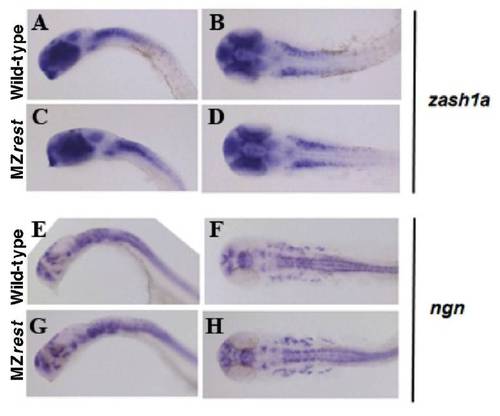
Neural progenitors are unaffected in rest mutants. WISH of proneural markers zash1a (A-D) and ngn (E-H) in 24-hpf wild type and MZrestsbu29/sbu29 mutants. (A,D,E,G) Lateral views; (B,D,F,H) dorsal views; anterior to the left. Examination of proneural markers zash1a and ngn did not reveal any differences between stage-matched wild type and rest mutants. |
|
Rest protein is expressed in FBMNs. Dorsal views of 48-hpf embryos, anterior to left. (A) Immunohistochemistry of wild-type embryos reveals Rest localization in reticulospinal neurons (white arrowhead), including the Mauthner (M) neuron (arrow), FBMNs (bracket), vagal motor neuron nuclei (black arrowhead), as well as in the surrounding neuroepithelium. (B) Staining of restsbu29/sbu29 homozygous mutant embryos does not detect Rest protein. Controls lacking the primary Rest antibody showed similar background fluorescence. |

Unillustrated author statements |