- Title
-
Mosaic Eyes is a novel component of the Crumbs complex and negatively regulates photoreceptor apical size
- Authors
- Hsu, Y.C., Willoughby, J.J., Christensen, A.K., and Jensen, A.M.
- Source
- Full text @ Development
|
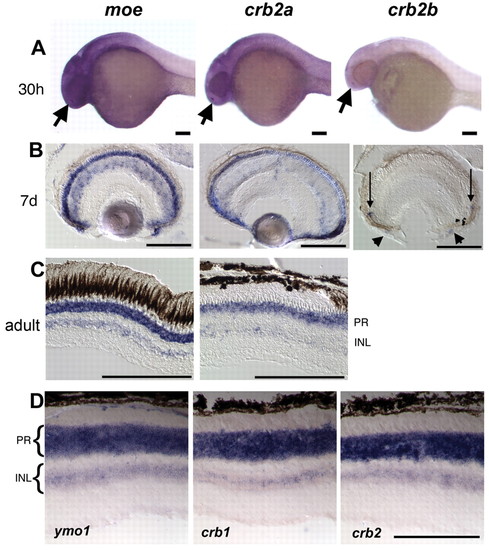
Expression of zebrafish crb2 genes. (A) moe and crb2a are expressed in the brain, eye and nose (arrows) at 30 hpf; crb2b is expressed in the nose (arrow) and weakly or not at all in the brain and eye. (B) moe and crb2a are expressed in photoreceptors, cells in the inner nuclear layer, progenitor cells in the periphery and cells in the anterior chamber at 7 dpf; crb2b expression is detected in a small number of newly differentiating photoreceptors near the ciliary margin (arrows) and in cells in the anterior chamber (arrowheads). (C) Zebrafish moe and crb2a are expressed in photoreceptors and cells in the inner nuclear layer in adults. (D) Mouse ymo1, crb1 and crb2 are expressed in photoreceptors and cells in the inner nuclear layer in the adult. Scale bars: 100 μm. INL, inner nuclear layer; PR, photoreceptors. |
|
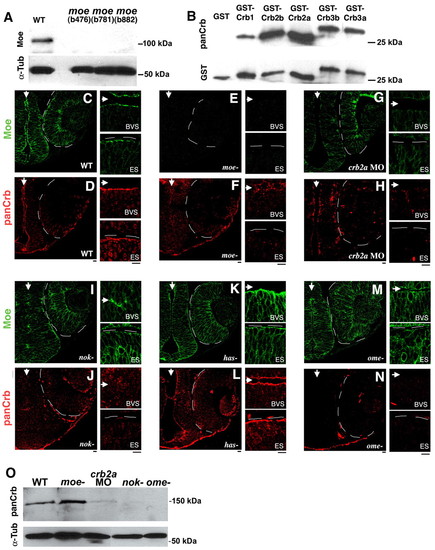
Localization of Moe and Crumbs proteins in embryonic brain and eye. (A) Western blot of wild type, moeb476, moeb781, moeb882 3d larvae probed with rabbit anti-Moe. Anti-Moe recognizes a protein of approximately 110 kDa in wild-type extracts that is absent in moe mutants. The blot was reprobed with anti-α-Tubulin as a loading control. (B) Anti-CRB3 recognizes the intracellular domains of all zebrafish Crumbs proteins. Western blot of GST, GST-Crb1intra, GST-Crb2aintra, GST-Crb2bintra, GST-Crb3aintra, GST-Crb3bintra probed with anti-CRB3 antibodies. The blot was reprobed with anti-GST as a loading control. (C-N) Moe (green) and panCrb (red) labeling in 30 hpf brain and eye in wild-type (C,D), moeb781 (E,F), crb2a morphant (G,H), nok (I,J), has (K,L) and ome (M,N) embryos. Arrows indicate brain ventricles and eyes are outlined. High magnification of the apical ventricular surface of the brain (BVS) and eye/retinal surface (ES) are shown to the right. Scale bars: 10 μm. (O) Western blot of 30 hpf wild-type, moeb781, crb2a morphant, nok and ome embryos probed with anti-panCrb antibodies. One-fifth of the material loaded for the panCrb Western was probed with anti-α-Tubulin to show relative loading amounts. EXPRESSION / LABELING:
|
|
Localization of Moe, panCrb, Nok and aPKCλ in the retina. (A,B) Colocalization of guinea pig anti-Moe and panCrb labeling in the peripheral region in 7 dpf eye (A) and photoreceptor layer at 4 dpf (B). AC, anterior chamber; PC, proliferating cells; PR, photoreceptors. (C,D) Colocalization of guinea pig anti-Moe and anti-Nok labeling in the peripheral region in 7 dpf eye (C) and photoreceptor layer at 4 dpf (D). (E,F) Colocalization of guinea pig anti-Moe and anti-aPKCλ labeling in the peripheral region in 7 dpf eye (E) and photoreceptor layer at 4 dpf (F). Insets are magnification of boxed areas. Scale bars: 20 μm; 10 μm in insets. |
|
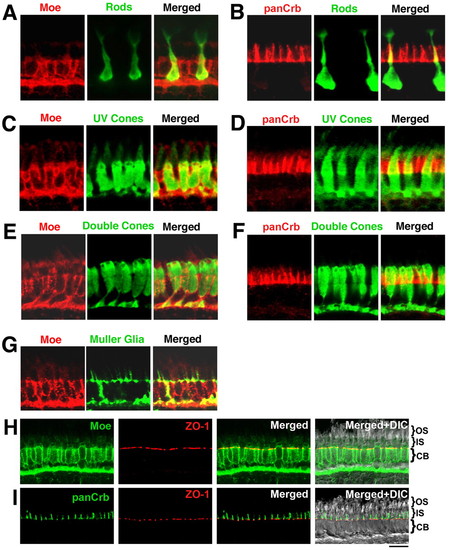
Localization of Moe and panCrb in specific retinal cell types at 5 dpf. (A) Merged guinea pig anti-Moe labeling of rod photoreceptors (in transgenic Xop-GFP fish) with rabbit anti-GFP. (B) Merged rabbit anti-panCrb labeling of GFP+ rod photoreceptors with GFP (in transgenic Xop-GFP fish). (C) Merged guinea pig anti-Moe labeling of UV cones with GFP (in transgenic UV-GFP fish) labeled with mouse anti-GFP. (D) Merged rabbit anti-panCrb labeling of UV cones with GFP (in transgenic UV-GFP fish) labeled with mouse anti-GFP. (E) Merged rabbit anti-Moe labeling of double cones with mouse Zpr-1 antibodies. (F) Merged rabbit anti-panCrb labeling of double cones with mouse Zpr-1 antibodies. (G) Merged guinea pig anti-Moe labeling of Müller glial cells with rabbit anti-Carbonic anhydrase II antibodies. (H) Rabbit anti-Moe, mouse anti-ZO-1, merged anti-Moe and anti-ZO-1, and merged with corresponding DIC image. (I) Rabbit anti-panCrb, mouse anti-ZO-1, merged anti-Moe and anti-ZO-1, and merged with corresponding DIC image. Scale bar: 10 μm. CB, cell bodies. EXPRESSION / LABELING:
|
|
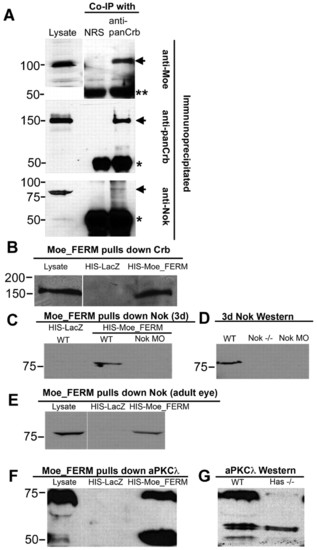
Moe forms a complex with Crumbs proteins, Nok (Pals1), and Has (aPKC&lambda). (A) Anti-panCrb (anti-CRB3) antibodies coimmunoprecipitate Moe (recognized by guinea pig anti-Moe), Crb2a/b, and Nok from adult eye. NRS, normal rabbit serum. Arrows indicate Moe, Crb2a/b, Nok protein. One asterisk indicates rabbit antibody; two asterisks indicate cross-reactivity of anti-guinea pig-HRP with rabbit antibody. (B) His-Moe_FERM, but not His-LacZ, pulls down a ∼150 kDa protein from 3 dpf larval lysates that is recognized by anti-panCrb antibodies. (C) His-Moe_FERM, but not His-LacZ, pulls down an ∼80 kDa protein from wild-type 3 dpf larval lysates, but not from nok morphant lysates, recognized by anti-Nok antibodies. (D) Anti-Nok antibodies recognize an ∼80 kDa protein in 3 dpf wild-type lysates that is absent in nok mutants and morphants in western blot. (E) His-Moe_FERM, but not His-LacZ, pulls down an ∼80 kDa protein from adult eye lysates recognized by anti-Nok antibodies. (F) His-Moe_FERM pulls down ∼75 kDa and ∼50 kDa proteins from 3 dpf larvae lysates that are recognized by anti-PKCλ antibodies. (G) Anti-PKCλ antibodies recognize proteins of approximately 75, 57, 54 and 50 kDa in western blot of 3 dpf wild-type lysates; the 75 kDa and 50 kDa proteins are absent in lysates from has mutants (Has-/-). |
|
Moe directly binds Crumbs proteins and Nok (Pals1). (A) His-Moe_FERM binds GST-Crb1, GST-Crb2a and GST-Crb2b. The blot was probed with anti-GST antibodies. Loading control, anti-HIS western blot of one-fortieth the material used in the experiment. (B) Far western analysis shows that membrane immobilized His-Moe_FERM (molecular weight approximately 55 kDa) binds GST-Crb1, GST-Crb2a and GST-Crb2b, but not GST alone, and no GST immunoreactivity is observed when His-Moe_FERM is incubated with buffer alone. Far westerns were probed with anti-GST antibodies. (C) GST-Nok-Int binds MBP-Moe_FERM but not MBP-Moe_C Term. The blot was probed with anti-MBP. Loading control, anti-GST western blot of one-twentieth the amount of MBP proteins loaded. (D) MBP-Moe_FERM but not MBP-Moe_C Term binds to membrane immobilized His-Nok_FL (Full Length) but not to His-Nok_N-term, which does not contain the predicted Band 4.1-binding domain. Far Westerns probed with anti-MBP. Loading control, anti-HIS Western blot of one-tenth the amount of His-fusion proteins. |
|
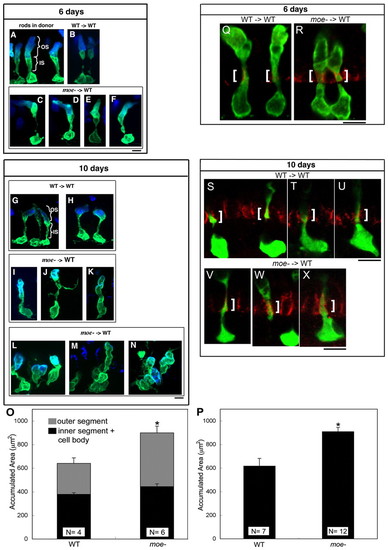
Moe is required for normal photoreceptor morphology and negatively regulates apical size. (A,B) Wild-type GFP+ rods labeled with anti-Rhodopsin antibody (blue) in a wild-type donor retina (A) and wild-type GFP+ rods transplanted into a wild-type host (B) at 6 dpf. (C-F) moe- GFP+ rods transplanted into wild-type hosts at 6 dpf. (G,H) Wild-type GFP+ rods transplanted into wild-type hosts at 10 dpf. (I-K) moe- GFP+ rods transplanted into wild-type hosts with few moe- neighbors at 10 dpf. (L-N) moe- GFP+ rods transplanted into wild-type hosts with large numbers of moe- neighbors at 10 dpf. (L) Three rods, (M) two to three intermingled rods and (N) three intermingled rods. (O) At 6 dpf, the accumulated area moe- rods is larger than wild-type rods. The increase in size is accounted for largely by an increase in outer segments whereas there is no significanct difference in the inner segments plus cell body (error bars shown; s.e.m.; *, P=0.04). The total accumulated area of moe- rods (901+/-64) is significantly greater than wild-type rods (641+/-37) (P=0.015). Numbers of individuals (N) included. (P) At 10 dpf, the accumulated area of moe- rods is larger than wild-type rods. (*, P=0.001; Student's t-test). (Q-X) panCrb labeling (red) in a single optical section (0.38 μm) of wild-type (Q) and moe- (R) GFP+ rods at 6 dpf and wild-type (S-U) and moe- GFP+ rods (V-X) at 10 dpf. White brackets indicate the region of panCrb localization in the inner segment. Scale bars: 5 μm. |
|
Injection of crb2a splice-blocking morpholinos phenocopies the moe mutations. (A) crb2a splicing, measured by RT-PCR, is reduced by injection of splice-blocking morpholino antisense oligonucleotides. Morpholino injection results in the amplification of a 1097 bp fragment. The reactions were also performed with no reverse transcriptase (no RT). (B-D) Head morphology of 30-hour wild-type (B), moeb476 mutant (C) and crb2a morphant (D) embryos. (E-G) The retinal pigmented epithelium is uniform in wild type (E) and patchy in moeb781 mutants (F) and crb2a morpholinos-injected embryos (G) at 48 hours. (H-J) The three nuclear layers are clearly defined in the wild-type retina (H), but are not defined in the moeb781 mutant (I) and the crb2a morphant (J) at 3 days. (K-M) Retinal ganglion cells (labeled using Zn5 antibodies) localize to the innermost part of the retina in the wild-type retina (K), but are disorganized and scattered throughout the retina in moeb781 mutants (L) and crb2a morphants (M) at 3 days. (N-P) Rod photoreceptors (visualized by genetically-encoded GFP) localize to the outermost part of the retina in the wild-type retina (N), but are disorganized and scattered throughout the retina in moeb781 mutants (O) and crb2a morphants (P) at 4 days. Eyes are outlined. Scale bars: in D, 500 μm for B-D; in P, 100 μm for E-P. |
|
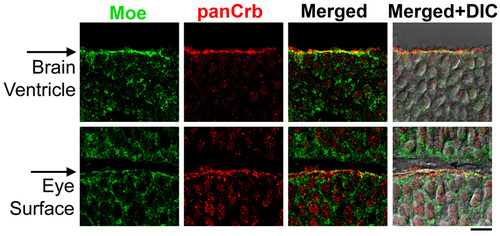
Co-localization of Moe and panCrb in the brain and eye at 30 hours. Moe is shown in green (guinea pig anti-Moe) and panCrb in red (rabbit anti-CRB3). Images are merged with corresponding DIC images. Apical surfaces are indicated with arrows. Scale bar: 10 μm. |
|
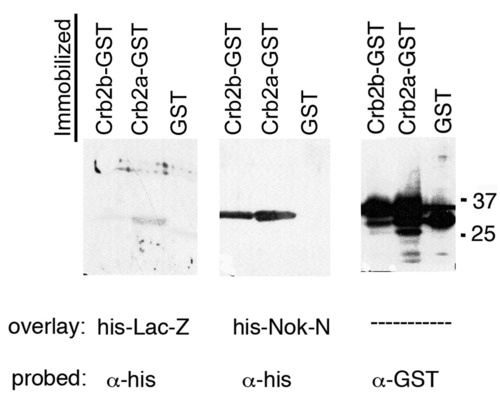
Nok (Pals1) interacts directly with zebrafish Crumbs proteins. His-Nok-N binds to membrane-immobilized GST-Crb2a and GST-Crb2b but not to GST alone in a far western probed with anti-His. His-LacZ does not show significant binding to any of the GST fusions. Control loading comprises anti-GST western blot with 1/50th amount of GST-fusion proteins. |
|
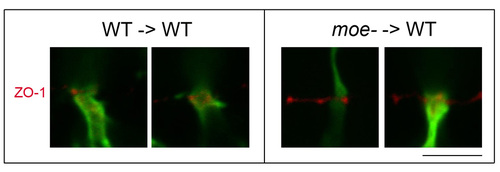
ZO-1 labeling of the outer limiting membrane in transplanted WT and moe- GFP+ rods at 6 days. ZO-1 is shown in red, GFP in green. Scale bar: 5 μm. |