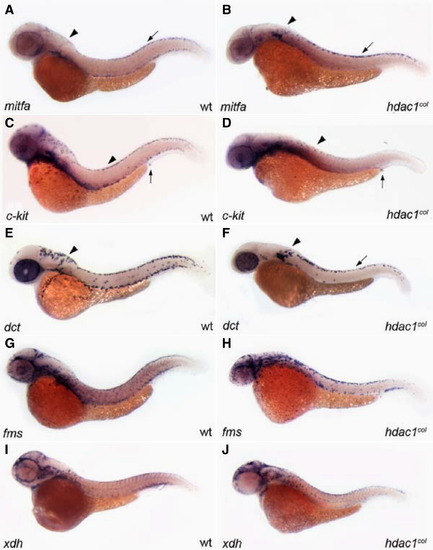
Melanophore development does not recover in hdac1col mutants. All panels are lateral views of 48 hpf embryos that are stained by in situ hybridization to reveal expression of mifta (A, B), c-kit (C, D), dct (E, F), fms (G, H) and xdh (I, J). (A, B) By 48 hpf mitfa expression is switched off in differentiating melanoblasts in wild-type; however in hdac1col mutants, mitfa continues to be expressed robustly in melanoblasts. (A, B, E, and F) There are still reduced numbers of mitfa and dct expressing melanoblasts in hdac1col mutants as compared to wild-type. Additionally, there is still a migration defect as most of the mitfa and dct expressing melanoblasts are located at their site of origin in the post otic region (arrowhead) and in the dorsal stripe (arrow). (C, D) In contrast to levels of mitfa and dct expression in melanoblasts in hdac1col mutants, c-kit expression in melanoblasts does not recover to wild-type levels (arrowheads). (G–J) Unlike melanophore development, xanthophore number and migration recover by 48 hpf, where there are numerous migrating fms-positive and xdh-positive xanthoblasts in wild-type and hdac1col mutants.
|

