- Title
-
Differential expression of enzymes in thymidylate biosynthesis in zebrafish at different developmental stages: implications for dtymk mutation-caused neurodegenerative disorders
- Authors
- Frisk, J.H., Örn, S., Pejler, G., Eriksson, S., Wang, L.
- Source
- Full text @ BMC Neurosci.
|
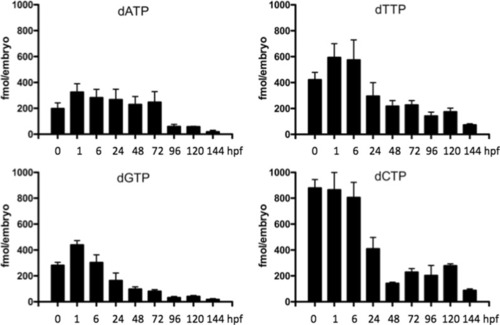
The levels of dNTPs at different developmental stages. A total of 50–75 embryos/larvae were collected at each time point and used in dNTP measurement (see Materials and Methods for details). Data were from 9 to 12 independent measurements and given as fmol dNTP per embryo (mean ± SD) |
|
The levels of TMPK at different developmental stages. TMPK activity in the cytosol ( |
|
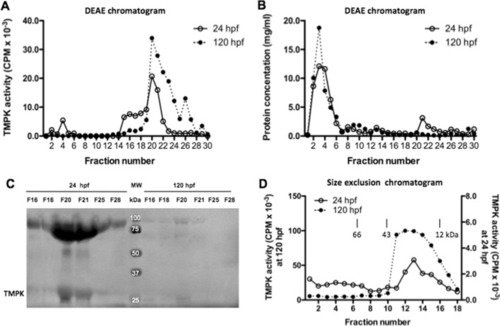
Characterization of cytosolic TMPK. A Partial purification of zebrafish TMPK by DEAE chromatography using cytosolic proteins isolated from zebrafish embryos harvested at 24 hpf and 120 hpf. The bound proteins were eluted with stepwise increase of KCl concentration (from 50 to 250 mM) in DEAE buffer. Fractions were collected and assayed for TMPK activity; B Protein concentration of the corresponding DEAE fractions; C Identification of the canonical TMPK proteins in the TMPK activity peak fractions by western blot analysis using a zebrafish Dtymk-specific antibody. Fractions F16, F18, F20, F21, F25, and F28 from DEAE chromatography were used and only part of the image is shown; D Size exclusion chromatography of partially purified zebrafish TMPK. Fractions F20 and F21 from the DEAE chromatography were used. Fractions were collected and assayed for TMPK activity. The Y-axis on the left side corresponded the TMPK activities from 120 hpf embryos and the Y-axis on the right side corresponded the TMPK activities from 24 hpf embryos. The elution positions of molecular weight markers are indicated |
|
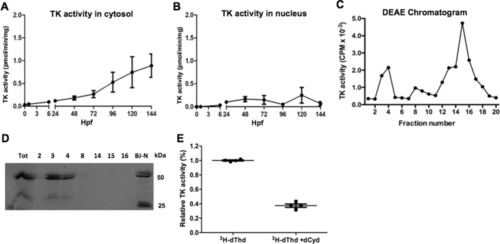
Characterization of thymidine kinases. TK activity in the cytosol (A) and nucleus (B) fractions at different zebrafish developmental stages. C DEAE chromatography of cytosolic protein isolated from zebrafish at 120 hpf. Fractions were collected and TK activity was determined by using [3 H]-dThd as substrate. D Western blot analysis of the TK peak fractions (2, 3, 4, 8, 14,15 and 16) from DEAE chromatography using a TK1 specific antibody and only part of the image is shown. Cell extracts from a fibroblast (BJ-N) cell line was used as control. E Fractions from DEAE chromatography (fraction 15 and 16) were assayed in the absence and presence of excess dCyd using [3 H]-dThd as the substrate. Data was plotted as mean ± SD |
|
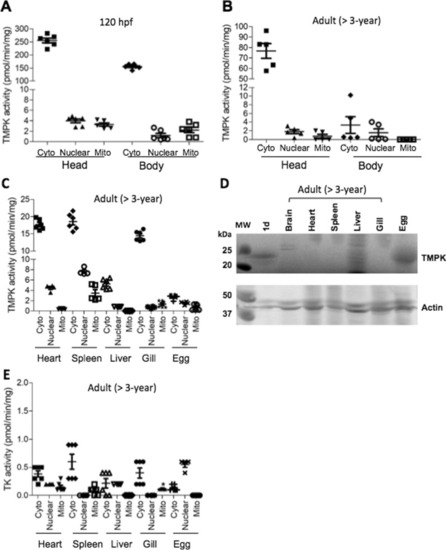
Organ distribution of TMPK and TK. TMPK activity in larvae at 120 hpf (A) and adult fish (> 3 years old) (B and C). D TMPK protein levels in different organs. E TK activity in different organs. The head and truncated body (without head) as well as organs from adult fish were used to isolate cytosolic, mitochondrial and nuclear fractions. Total proteins were extracted and used to measure TMPK and TK activity by using [3 H]-labeled substrates and the data was plotted as mean ± SD. Western blot analysis was conducted by using the zebrafish Dtymk-specific antibody and only parts of the images are shown |
|
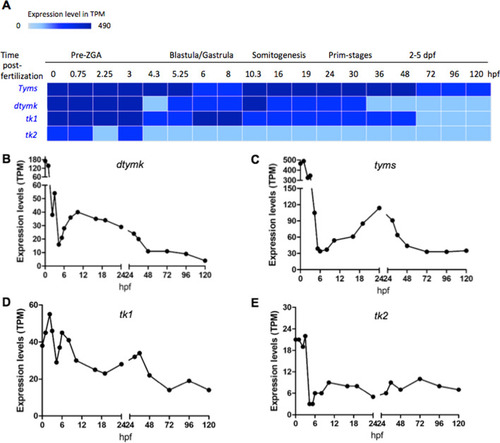
mRNA expression profiles of selected enzymes involved in dTTP synthesis at different zebrafish developmental stages. A mRNA expression heatmap. dtymk, thymidylate kinase; tyms, thymidylate synthesis; tk1, thymidine kinase 1; tk2, thymidine kinase 2. Plots of mRNA levels of dtymk (B), tyms (C), tk1 (D) and tk2 (E). TPM, transcript per million. Data are shown as average expression (mean TPM) and retrieved from Expression atlas – baseline expression of transcriptional profiling of zebrafish developmental stages (https://www.ebi.ac.uk/gxa/experiments/E-ERAD-475/Result) |