Fig. 6
- ID
- ZDB-FIG-220607-49
- Publication
- Sánchez-Iranzo et al., 2022 - Strength of interactions in the Notch gene regulatory network determines patterning and fate in the notochord
- Other Figures
- All Figure Page
- Back to All Figure Page
|
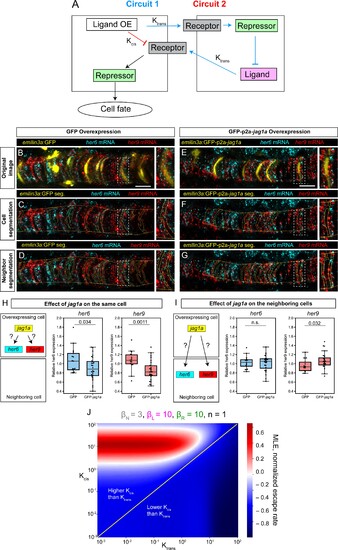
(A) Two possible circuits may explain the effect of jag1a on fate of the cell where jag1a is overexpressed. Circuit 1 is based on the interaction of ligand and receptor in trans. Circuit 2 is based on a possible role of cis-inhibition of the Notch receptor by the ligand. Cells where we overexpress the ligand are represented as the cell on the left. Adjacent cells are represented on the right. OE, overexpression. (B – G), Airyscan confocal planes of fixed 22 hpf transgenic fish injected with emilin3a:GFP (B–D) or emilin3a:GFP-p2a-jag1a (E–G) constructs. GFP was detected by antibody staining and her6 and her9 mRNA by in situ HCR in whole mount embryos. (C and F) show the notochord outline manually selected and the outline of GFP-positive cells automatically segmented. (D and G) show the outline of the manually selected notochord and the neighborhood to the GFP-positive cells. On the right side of each panel, a magnified view of the boxed region is shown. (H, I) Quantification of her6 and her9 mRNA expression after GFP-based segmentation as shown in (C, F) or (D, G), respectively. Values of her6 and her9 expression levels inside the segmented area inside the notochord were divided by the expression levels of the same genes in the region outside the segmented area, also inside the notochord. Each point represents a different fish. Two-tailed p-values are shown in the plots. n.s., non-significant. (J) Escape rates from the homogeneous steady state (indicated by Maximum Lyapunov Exponents, or MLE) as a function of Kcis and Ktrans parameters. Positive MLE values (red) support patterning, while negative MLE values (blue) do not. Scale bars, 20 μm. |