FIGURE
Figure 3
- ID
- ZDB-FIG-211120-120
- Publication
- Safarian et al., 2021 - Panx1b Modulates the Luminance Response and Direction of Locomotion in the Zebrafish
- Other Figures
- All Figure Page
- Back to All Figure Page
Figure 3
|
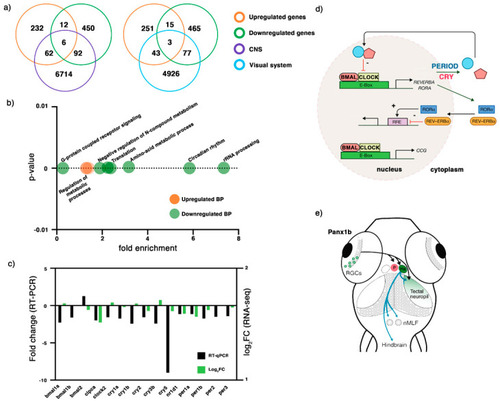
Figure 3. RNA-seq analysis of 6dpf panx1b−/− larvae. (a) Comparison of differentially regulated RNA transcripts against two categories: central nervous system (CNS) and the visual system. (b) GO annotation of RNA-seq data in terms of biological process (BP): the upregulated RNA transcripts were enriched in the regulation of metabolic processes, whereas the downregulated genes were annotated for RNA processing, circadian rhythms, amino-acid metabolic process, translation, negative regulation of nitrogen (N) compounds metabolism, and G-protein coupled receptor signaling. (c) Validation of RNA-seq data by RT-qPCR. The significant downregulation of cry1, cry5, bmal1a, bmal1b, and clock2 genes in 6 dpf panx1b−/− larvae was confirmed by RT-qPCR in n = 3 independent replicates. (d) Cartoon illustrating the network of circadian clock genes differently regulated in panx1b−/− larvae. Regulated genes in bold. (e) Illustration of the connection of the inner retina with optic tectum, habenula (Ha), and pretectum (p) conferring light sensitivity and circadian rhythmicity, and descending control of locomotion. Abbreviation: nMLF, nucleus of the medial longitudinal fasciculus.
|
Expression Data
| Genes: | |
|---|---|
| Fish: | |
| Anatomical Term: | |
| Stage: | Day 6 |
Expression Detail
Antibody Labeling
Phenotype Data
| Fish: | |
|---|---|
| Observed In: | |
| Stage: | Day 6 |
Phenotype Detail
Acknowledgments
This image is the copyrighted work of the attributed author or publisher, and
ZFIN has permission only to display this image to its users.
Additional permissions should be obtained from the applicable author or publisher of the image.
Full text @ Int. J. Mol. Sci.