Fig. 4
- ID
- ZDB-FIG-081104-7
- Publication
- Batista et al., 2008 - Pax2/8 act redundantly to specify glycinergic and GABAergic fates of multiple spinal interneurons
- Other Figures
- All Figure Page
- Back to All Figure Page
|
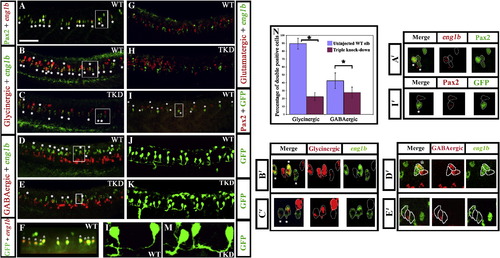
Pax2/8 specify inhibitory fates in CiA interneurons. Lateral views of 24 h trunks. (A, B, D and G) WT embryos, (C, E and H) triple-knock-down (TKD) embryos, (F, I, J and L) Tg(pax2a:GFP) WT embryos (K and M) Tg(pax2a:GFP) TKD embryos. Rostal is left, dorsal is top. (A) Pax2 (green), eng1b (red). (B–E, G and H) Double in situ hybridisation for eng1b (green) and neurotransmitter markers (red). eng1b staining is weaker in some pictures, but the same number of eng1b-expressing cells (13.9 ± 1.8 over a 5-somite length) were counted in each embryo (control and TKD). (F) Eng1b (red), GFP (green). The vast majority of GFP-positive cells in Tg (pax2a:GFP) embryos are CiAs and express Eng1b, although there are also a few more dorsal GFP-positive cells that are not CiAs. Occasionally a weak Eng1b-expressing cell can be observed that is not yet expressing GFP. (I) Pax2 (red), GFP (green). Note that all green cells are also red but several red cells are not green. (J–M) GFP (green). All of the ventral GFP-expressing cells are CiAs with similar ipsilateral ascending axons. (L and M) show higher magnification views of individual GFP-expressing CiAs in Tg(pax2a:GFP) WT (L) and TKD (M) embryos. For quantification of soma size and axon length see Supp. Data Table 3. Stars indicate double labelled cells. Scale bar = 50 μm (A–E and G–I); 30 μm (F, J and K). (N) Percentage of CiAs (eng1b-expressing cells) that are glycinergic or GABAergic in WT and TKD embryos. Each percentage is an average from 12 different embryos. Stars indicate statistically significant results (p < 0.05). Error bars denote standard deviation. No glutamatergic CiAs were observed in any embryo. (A′–E′ and I′) are magnified views of boxes in A–E and I respectively, showing red and green channels and merged images for a single confocal focal plane. Stars in merged images indicate double labelled cells. For single channel images of the whole lateral view for A–E and I see Supp. Data Fig. 6. |
| Genes: | |
|---|---|
| Antibody: | |
| Fish: | |
| Knockdown Reagents: | |
| Anatomical Term: | |
| Stage: | Prim-5 |
Reprinted from Developmental Biology, 323(1), Batista, M.F., and Lewis, K.E., Pax2/8 act redundantly to specify glycinergic and GABAergic fates of multiple spinal interneurons, 88-97, Copyright (2008) with permission from Elsevier. Full text @ Dev. Biol.