- Title
-
The Hox cofactors Meis1 and Pbx act upstream of gata1 to regulate primitive hematopoiesis
- Authors
- Pillay, L.M., Forrester, A.M., Erickson, T., Berman, J.N., and Waskiewicz, A.J.
- Source
- Full text @ Dev. Biol.
|
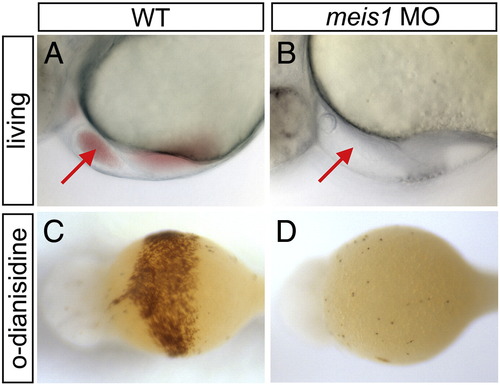
meis1-morphant embryos fail to produce visible circulating erythrocytes. (A, B) Lateral view of live 48 hpf embryos showing close-up of heart region; anterior to the left. Unlike their wild type (WT) counterparts (A), meis1-morphant (B) embryos lack visible circulating blood cells. Arrows indicate heart. (C, D) o-dianisidine staining of differentiated erythrocytes in 48 hpf whole-mount embryos; ventral view with anterior to the left. o-dianisidine staining is abolished in meis1-morphant (D) embryos when compared to WT embryos (C). |
|
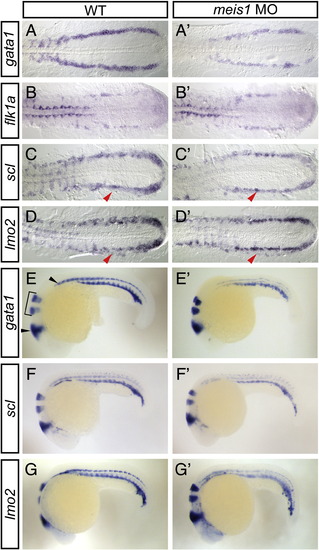
meis1-morphant embryos exhibit defects in primitive hematopoietic gene expression. Shown are representative embryos following in situ hybridization analysis of hematopoietic marker expression in wild type (WT; A–G) compared with meis1-morphant (A′–G′) embryos. (A–D′) The PLM of 16 hpf flat-mounted, deyolked embryos is shown in dorsal view with anterior to the left. gata1 expression is severely reduced in meis1-morphant embryos (A′; 85%, n = 39) when compared to WT (A). flk1a expression is unchanged in meis1-morphant embryos (B′; 100%, n = 21) when compared to WT (B). Lateral domain of scl (90%, n = 29) and lmo2 expression (79%, n = 24) is abolished in meis1-morphant embryos (C′, D′; red arrowheads), while the medial domain of scl and lmo2 expression is near normal. (E–G′) 24 hpf whole-mount embryos are shown in lateral view with anterior to the left. eng2a expression in the midbrain hindbrain boundary and muscle pioneers (E; black arrowheads) and egr2b expression in hindbrain rhombomeres 3 and 5 (E; bracket) is shown in all panels. gata1 (E′; 91%, n = 53), scl (F′; 97%, n = 35), and lmo2 (G′; 92%, n = 36) expression is reduced in the ICM of meis1-morphant embryos when compared to WT (E–G). |
|
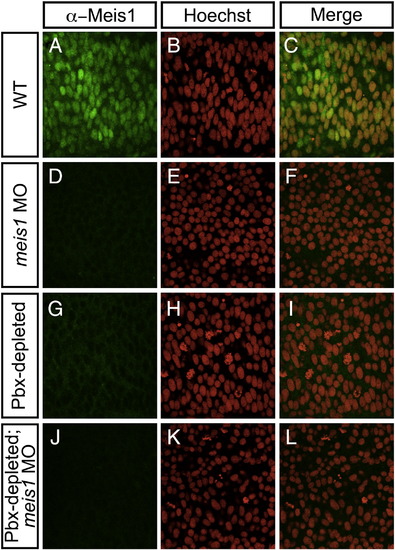
Meis1 protein levels are severely diminished in the posterior mesoderm of Pbx-depleted and meis1-morphant embryos. Shown are representative embryos following immunohistochemical staining with the P2A6 antibody (green; A, D, G, J) to visualize Meis1 protein levels, and Hoechst 33258 (red; B, E, H, K) to visualize nuclei in 14 hpf wild type (WT; A–C), meis1-morphant (D–F), Pbx-depleted (G–I), and Pbx-depleted; meis1-morphant (J–L) embryos. Flat-mount; dorsal view of posterior mesoderm; anterior to left. All embryos visualized under a 100x objective. In wild type (WT; A–C) embryos, α-Meis1 antibody staining is punctate, and colocalizes with Hoechst nuclear staining. meis1-morphant (D) and Pbx-depleted (G) embryos exhibit severely decreased α-Meis1 antibody staining when compared to WT embryos (A). α-Meis1 antibody staining is further abolished in Pbx-depleted; meis1-morphant embryos (J). α-Meis1 antibody staining does not colocalize with Hoechst nuclear staining in Pbx-depleted embryos (G–I). |
|
Pbx and Meis1 act in a cooperative fashion to regulate erythroid gene expression. (A–H) Shown are representative embryos following in situ hybridization analysis of gata1 (A–D) and hbae3 (E–H) expression in 16 hpf embryos. Embryos shown are deyolked and flat-mounted with a dorsal view of gene expression in the PLM and anterior oriented to the left. Embryos partially depleted of Pbx (injected with 4.5 ng of pbx2/4 MO) exhibit subtle decreases in gata1 (B; 83%, n = 18) and hbae3 (F; 71%, n = 7) expression. Embryos partially depleted of Meis1 (injected with 2 ng of meis1 MO) exhibit subtle decreases in gata1 (C; 85%, n = 20) and hbae3 (G; 100%, n = 6) expression. Embryos partially depleted of both Pbx and Meis1 exhibit profoundly reduced gata1 (D; 90%, n = 29) and hbae3 (H; 95%, n = 19) expression. (I) Quantitative real-time PCR analysis of gata1 expression in 16 hpf wild type embryos (WT), lzr/pbx4+/- incross progeny injected with a full dose of pbx2/4 MO (pbx2/4 MO), meis1-morphant embryos (meis1 MO), and lzr/pbx4+/- incross progeny injected with a full dose of both pbx2/4 MO and meis1 MO (pbx2/4 MO; meis1 MO). Shown is the relative quantity of gata1 expression. Samples were normalized to ef1a and WT was set to 1. Error bars indicate standard deviation from the mean. *Indicates the difference compared with WT is significant by Student t test, P < 0.0001. **Indicates the difference compared with all other samples is significant by Student t test, P < 0.0001. |
|
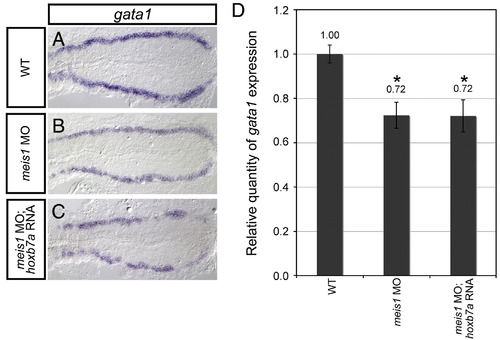
Overexpressing hoxb7a rescues gata1 erythroid gene expression in Pbx-depleted but not meis1-morphant embryos. (A–C) Shown are representative embryos following in situ hybridization analysis of gata1 expression in 16 hpf embryos. Dorsal view of PLM gene expression is shown in flat-mounted and deyolked embryos, with anterior oriented to the left. Both meis1-morphant embryos (B; 94%, n = 63) and hoxb7a RNA-injected meis1-morphant embryos (C; 91%, n = 43) exhibit a severe decrease in gata1 expression when compared to wild type embryos (WT; A). (D) Quantitative real-time PCR analysis of gata1 expression in 16 hpf wild type embryos (WT), meis1-morphant embryos (meis1 MO), and hoxb7a RNA-injected meis1-morphant embryos (meis1 MO; hoxb7a RNA). Shown is the relative quantity of gata1 expression. Samples were normalized to ef1a and WT was set to 1. Error bars indicate standard deviation from the mean. *Indicates the difference compared with WT is significant by Student t test; P < 0.0001. |
|
Pbx and Meis1 act in a cooperative fashion to regulate primitive myelopoietic gene expression. (A–D) Shown are representative embryos following in situ hybridization analysis of pu.1 expression in 24 hpf embryos. Dorsal view of gene expression in the anterior lateral-plate mesoderm (ALPM) is shown in whole-mount embryos with anterior oriented to the left. meis1-morphant (B), Pbx-depleted (C), and meis1-morphant; Pbx-depleted (D) embryos exhibit a severe increase in the number of pu.1-expressing cells when compared to wild type (WT; A) embryos. (E) Quantification of the phenotypes shown in A–D. Shown is the average number of pu.1-expressing cells in the ALPM of 24 hpf embryos as determined by in situ hybridization. Error bars indicate standard error of the mean. *Indicates the difference compared with WT is significant by Student t test; P < 0.0001. **Indicates the difference compared to all other samples is significant by Student t test; P < 0.001. (F–M) Shown are representative embryos following in situ hybridization analysis of pu.1 (F, G) in lateral view, and l-plastin (lcp1; H–J) and lysozyme C (lyz; K–M) expression in dorsal view 24 hpf whole-mount embryos. Pbx-depleted; meis1-morphant embryos (G) exhibit upregulated pu.1 expression in the intermediate cell mass when compared to WT embryos (F). Pbx-depleted (I, L), and meis1-morphant (J, M) embryos exhibit increased numbers of lcp1 and lyz-positive cells in the ALPM when compared to WT embryos (H, K). Genotype of embryos was determined by in situ hybridization analysis of egr2b (F–M) and eng2a (H–J) expression. |
|
Pbx and Meis1 act upstream of gata1 to regulate the expression of hbae3 and scl, but not lmo2. (A–H) Shown are representative embryos following in situ hybridization analyses of lmo2 and scl expression in lateral view 24 hpf whole-mount embryos. gata1-morphant embryos (B; 100%, n = 11) exhibit near normal lmo2 expression when compared to WT embryos (A). gata1-morphant (D; 100%, n = 7), meis1-morphant (E; 100%, n = 6), and Pbx-depleted (G; 94%, n = 16) embryos exhibit diminished scl expression in the ICM when compared to wild type (WT; C) embryos. meis1-morphant, gata1 RNA-injected embryos (F; 100%, n = 5), and Pbx-depleted, gata1 RNA-injected embryos (H; 95%, n = 19) exhibit greater levels of scl expression than respective meis1-morphant (E) and Pbx-depleted (G) counterparts. Genotype of embryos was determined by in situ hybridization analysis of egr2b expression in hindbrain rhombomeres 3 and 5 (A–H), and eng2a expression in the midbrain hindbrain boundary and muscle pioneers (C–H). (I–L) Shown are representative embryos following in situ hybridization analysis of hbae3 expression in 16 hpf embryos. Dorsal view of PLM gene expression is shown in deyolked, flat-mounted embryos with anterior oriented to the left. Pbx-depleted; meis1-morphant embryos (J; 90%, n = 10) exhibit nearly abolished hbae3 expression when compared to WT embryos (I). gata1 RNA-injected, Pbx-depleted; meis1-morphant embryos (K; 82%, n = 11) exhibit near normal hbae3 expression. gata1 RNA-injected embryos (L; 100%, n = 4) exhibit slightly increased hbae3 expression when compared to WT embryos (I). |
|
Unlike Cdx, Meis1 and Pbx function downstream of scl to activate gata1 expression. (A–H) Shown are representative embryos following in situ hybridization analysis of scl and gata1 expression in 12 hpf embryos. Dorsal view of gene expression in the posterior lateral-plate mesoderm (PLM) is shown in whole-mount embryos with anterior oriented to the left. Pbx-depleted; meis1-morphant embryos exhibit wild type (WT) levels of scl expression (B; 100%, n = 13) and abolished gata1 expression (D; 100%, n = 18) when compared to WT (A, C) embryos. Genotype of Pbx-depleted; meis1-morphant embryos was determined by in situ hybridization analysis of egr2b expression. Cdx-depleted embryos exhibit abolished scl (F; 100%, n = 5) and gata1 (H; 100%, n = 7) expression when compared to WT (E, G) embryos. (I) Hierarchical model indicating the genetic interactions that occur between a subset of transcription factors that regulate zebrafish primitive hematopoiesis. Arrows do not necessarily represent direct interactions. |
|
Lateral domain of scl expression colocalizes with pax2a expression. Shown are representative embryos following in situ hybridization analysis of scl and gata1 (A) or scl and pax2a (B) expression in wild type embryos. The PLM of 16 hpf flat-mounted, deyolked embryos is shown in dorsal view with anterior oriented to the left. Arrowheads indicate scl and pax2a colocalization. |
|
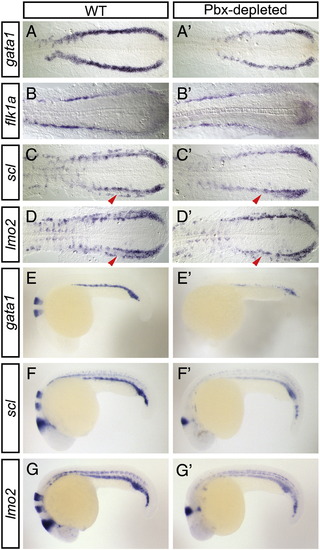
Pbx-depleted embryos exhibit defects in primitive hematopoietic gene expression. Shown are representative embryos following in situ hybridization analysis of hematopoietic marker expression in wild type (WT; A-G) compared with Pbx-depleted (A′-G′) embryos. (A-D′) The PLM of 16 hpf flat-mounted, deyolked embryos is shown in dorsal view with anterior to the left. gata1 expression is severely reduced in Pbx-depleted embryos (A′ 77%, n = 31) when compared to WT (A). flk1a expression is unchanged in Pbx-depleted embryos (B′ 100%, n = 31) when compared to WT (B). Lateral domain of scl (90%, n = 20) and lmo2 expression (90%, n = 31) is abolished in Pbx-depleted embryos (C′, D′ arrowheads), while the medial domain of scl and lmo2 expression is near normal. (E-G′) 24 hpf whole-mount embryos are shown in lateral view with anterior to the left. gata1 (E′ 88%, n = 66), scl (F′ 90%, n = 30), and lmo2 (G′ 94%, n = 32) expression is reduced in the ICM of Pbx-depleted embryos when compared to WT (E-G). |
|
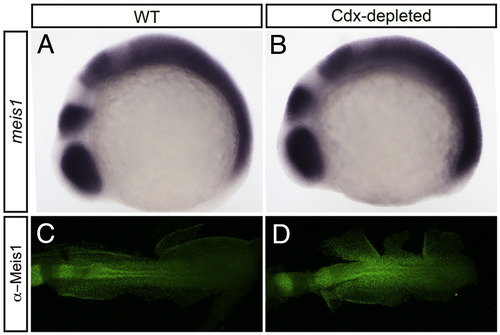
Cdx-depleted embryos exhibit increased levels of Meis1. (A, B) Shown are representative embryos following in situ hybridization analysis of meis1 expression in wild type (WT; A) compared with Cdx-depleted (B) embryos. The PLM of 14 hpf flat-mounted, deyolked embryos is shown in dorsal view with anterior to left. meis1 expression is increased in the posterior of Cdx-depleted embryos (B) when compared to WT (A). (C, D) Shown are representative embryos following immunohistochemical staining with the P2A6 antibody to visualize Meis1 protein levels in 14 hpf wild type (C) and Cdx-depleted (D) embryos. Flat-mount; dorsal view of embryo; anterior to left. Embryos are visualized under a 10x objective. Meis1 antibody staining is increased in Cdx-depleted embryos (D) when compared to WT (C). |
|
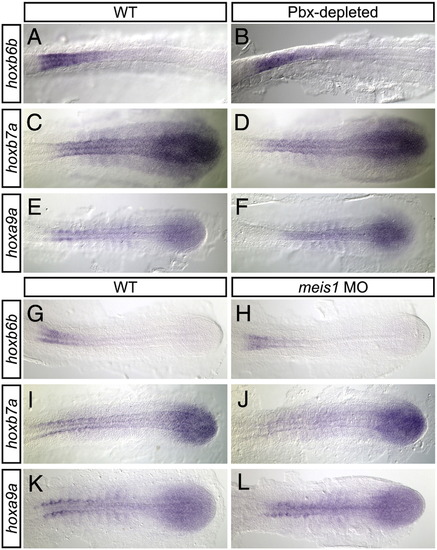
Pbx-depleted and meis1-morphant embryos exhibit normal posterior hox gene expression. Shown are representative embryos following in situ hybridization analyses of hoxb6b (A, B, G, H), hoxb7a (C, D, I, J), and hoxa9a (E, F, K, L) expression in wild type (WT; A, C, E, G, I, K), Pbx-depleted (B, D, F), and meis1-morphant (H, J, L) embryos. The PLM of 16 hpf flat-mounted, deyolked embryos is shown in dorsal view with anterior oriented to the left. In comparison to WT embryos, Pbx-depleted and meis1-morphant embryos exhibit normal hoxb6b, hoxb7a, and hoxa9a expression. |
|
Meis1 and Pbx are required for normal activation of gata1 expression. Shown are representative embryos following in situ hybridization analysis of gata1 expression in 12 hpf embryos. Dorsal view of gene expression in the posterior lateral-plate mesoderm (PLM) is shown in whole-mount embryos with anterior oriented to the left. Pbx-depleted (B) and meis1-morphant (C) embryos exhibit abolished gata1 expression when compared to wild type (WT; A). |
|
Overexpressing hoxb7a rescues gata1 erythroid gene expression in Cdx-depleted embryos, but not in Pbx-depleted; meis1-morphant embryos. Shown are representative embryos following in situ hybridization analysis of gata1 expression. (A–C) Dorsal view of PLM gene expression is shown in 16 hpf flat-mounted and deyolked embryos, with anterior oriented to the left. Both Pbx-depleted; meis1-morphant embryos (B) and hoxb7a RNA-injected Pbx-depleted; meis1-morphant embryos (C) exhibit a severe decrease in gata1 expression when compared to wild type embryos (WT; A). (D–F) Dorsal view of PLM gene expression is shown 12 hpf whole-mount embryos, with anterior oriented to the left. Cdx-depleted embryos (E) exhibit a notable decrease in gata1 expression when compared to WT embryos (D). Cdx-depleted, hoxb7a RNA-injected embryos (F) exhibit greater levels of gata1 expression than Cdx-depleted embryos (E). |
Reprinted from Developmental Biology, 340(2), Pillay, L.M., Forrester, A.M., Erickson, T., Berman, J.N., and Waskiewicz, A.J., The Hox cofactors Meis1 and Pbx act upstream of gata1 to regulate primitive hematopoiesis, 306-317, Copyright (2010) with permission from Elsevier. Full text @ Dev. Biol.