- Title
-
Systematic studies of all PIH proteins in zebrafish reveal their distinct roles in axonemal dynein assembly
- Authors
- Yamaguchi, H., Oda, T., Kikkawa, M., Takeda, H.
- Source
- Full text @ Elife
|
(A) Phylogenetic tree of PIH proteins in yeast, Chlamydomonas, and vertebrates. Amino acid sequences were aligned by MAFFT program with FFT-NS-i option (Katoh and Standley, 2013) and evolutionary distances were calculated using neighbor-joining (Saitou and Nei, 1987). yeast, Saccharomyces cerevisiae; Chlamy, Chlamydomonas reinhardtii; z, zebrafish (Danio rerio); o, medaka (Oryzias latipes); m, Mus musculus; h, Homo sapiens. (B) Genomic organization of PIH genes in zebrafish. Black: exons. Gray: untranslated regions. Red asterisks indicate the target sites of our genome-editing. (C) Immunoblot of PIH proteins. None of the PIH proteins were detected in the sperm flagella fraction (asterisks). ?-tubulin: control. (D) Whole-mount in situ hybridization of PIH genes. Arrows in lateral views indicate expression of PIH genes in ciliated organs (Kupffer?s vesicle, floor plate, otic vesicle, and pronephric duct). Flat mount preparations show dorsal views of the posterior regions of 12 hpf embryos. pih1d1 was ubiquitously expressed in 12 hpf embryos (black asterisk) containing Kupffer?s vesicle and floor plate (red arrowheads). ktu was also expressed in brain rudiments at 32 hpf (black arrowhead). White asterisks: non-specific staining of yolk. Scale bars: 200 ?m. (E and F) Diagrams of zebrafish embryos at (E) 12 hpf and (F) 32 hpf, showing typical ciliated organs. (G) Left: preparation procedure of flat mount. Right: diagram of Kupffer?s vesicle and floor plate in flat-mounted embryos. EXPRESSION / LABELING:
|

ZFIN is incorporating published figure images and captions as part of an ongoing project. Figures from some publications have not yet been curated, or are not available for display because of copyright restrictions. PHENOTYPE:
|
|
Cryo-ET revealed structural defects of axonemal dyneins in mutant spermatozoa. (A) DMT structure of native zebrafish sperm. Left: side view. Right: base-to-tip view. A-tub and B-tub: A- and B-tubule of DMT, respectively. (B) Diagram of the DMT structure of zebrafish sperm. f IC/LC means IAD f intermediate chain and light chain complex. (C) DMT structure of human respiratory cilia (EMD-5950; Lin et al., 2014). (D) DMT structure of Chlamydomonas flagella (DMT 2?8 averaged; EMD-2132; Bui et al., 2012). Red arrow indicates a linker between N-DRC and OAD. (E) Diagram of the DMT structure of Chlamydomonas flagella. (F?K) DMT structures of PIH gene mutant spermatozoa. For pih1d2-/-;ktu-/-, J (+OAD) and K (-OAD) represent subtomograms of axonemes with or without OADs, respectively. Red circles indicate the defects of axonemal dyneins. Green, OADs; red, IAD f; orange, IAD a; yellow, IAD b; light-green, IAD c; cyan, IAD e; indigo, IAD g; violet, IAD d; blue, RSs. |
|
Immunoblot and immunofluorescence microscopy of axonemal dyneins. (A) Immunoblot of sperm axonemes. Asterisks, filled circles, and open circles indicate missing, decreased, and shifted bands, respectively. Acetylated tubulin: control. (B) Immunofluorescence microscopy of zebrafish spermatozoa. Dnah8 was localized along the entire length of sperm flagella in WT, pih1d1-/-, pih1d2-/-, ktu-/-. In twister-/-, Dnah8 was lost (white asterisk), while in pih1d2-/-;ktu-/-, Dnah8 was localized only in the proximal half of the flagella (white arrowhead). |
|
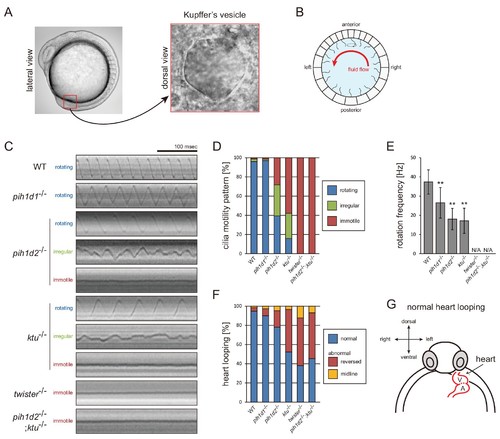
Mutations of PIH genes caused abnormal motilities of Kupffer?s vesicle cilia. (A) Images of zebrafish embryo and Kupffer's vesicle. Red square in left-hand image indicates the position of the Kupffer's vesicle in a 12 hpf embryo. Right-hand image shows the dorsal view of the Kupffer's vesicle. (B) Diagram of zebrafish Kupffer's vesicle. Mono-cilia, which project from epithelial cells, have rotational motilities to produce left-ward fluid flow (red arrow) in the organ. (C) Typical kymographs of Kupffer?s vesicle cilia in WT and PIH gene mutants. Patterns of the kymographs were categorized into three classes: rotating (blue), irregular (green), and immotile (red). (D) Ratios of each motility class. Number of observed cilia: WT, 76; pih1d1-/-, 67; pih1d2-/-, 198; ktu-/-, 190; twister-/-, 53; and pih1d2-/-;ktu-/-, 48. (E) Rotational frequencies of Kupffer?s vesicle cilia. Bar graphs show mean �SD. **: p-value<0.01 in Dunnett's test of each mutant against WT. (F) Heart looping of WT and mutant embryos at 30 hpf. Number of samples: WT, 247; pih1d1-/-, 288; pih1d2-/-, 275; ktu-/-, 285; twister-/-, 139; and pih1d2-/-;ktu-/-, 276. (G) Diagram represents normal heart looping of the 30 hpf embryo. V, ventricle; A, atrium. PHENOTYPE:
|
|
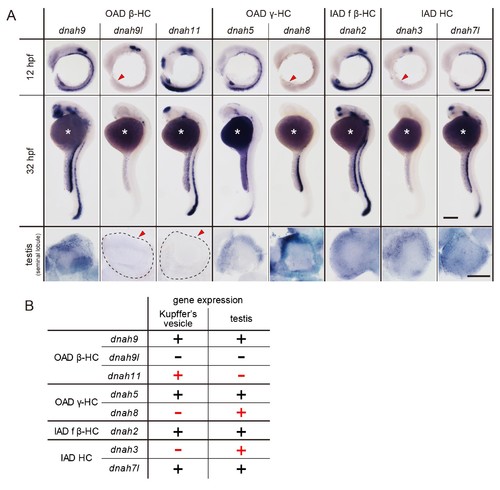
DNAH genes showed distinct expression patterns in zebrafish embryos and testis. (A) Whole-mount in situ hybridization of DNAH genes with embryos (12 and 32 hpf) and dissected testis (seminal lobule). For embryos, lateral views are shown. Yolk of 12 hpf embryos was removed before observation to show Kupffer?s vesicle clearly. Red arrowheads indicate Kupffer?s vesicles or testes in which DNAH gene expressions were not detected. White asterisks: non-specific staining of yolk. Scale bars: 200 ?m for embryos; 100 ?m for testes. (B) Comparison of DNAH gene expression between Kupffer?s vesicle and testis. +, expressed; -, expression not detected. Red indicates when DNAH gene expression was difference between the two organs. EXPRESSION / LABELING:
|
|
Expansion of pronephric duct was observed in twister-/- and pih1d2-/-;ktu-/-. (A?F) Dorsal views of zebrafish larvae at 4 days post-fertilization. (A) WT, (B) pih1d1-/-, (C) pih1d2-/-, (D) ktu-/-, (E) twister-/-, (F) pih1d2-/-;ktu-/-. twister-/- and pih1d2-/-;ktu-/- exhibited abnormal expansions of pronephric ducts (red arrowheads), which disabled larvae from having their pectoral fins close to their bodies. |