- Title
-
Active nuclear transcriptome analysis reveals inflammasome-dependent mechanism for early neutrophil response to Mycobacterium marinum
- Authors
- Kenyon, A., Gavriouchkina, D., Zorman, J., Napolitani, G., Cerundolo, V., Sauka-Spengler, T.
- Source
- Full text @ Sci. Rep.
|
Binary transgenic zebrafish model for regulatory profiling of neutrophils in response to M. marinum. (A) Driver line expressing BirA in neutrophils (i) is crossed with effector line ubiquitously expressing Avi-tagged Rangap (ii) for biotinylation of nuclei. In double transgenic fish, only the nuclei of neutrophils will be biotinylated (iii). (B) Effector line with ubiquitously expressed Avi-Cerulean-Rangap. (C) Neutrophil-specific BirA driver line with Citrine reporter. (D) Time course analysis of neutrophil infiltration of M. marinum. Embryos were injected with M. marinum at the 32-512-cell stage, fixed and immunofluorescence carried out for mpx protein at 2 dpi (i), 3 dpi (ii), 4 dpi (iii) and 5 dpi (iv). Neutrophil localisation is indicated by white arrows. (E) Neutrophil numbers in proximity to M. marinum were quantified at 2–5 dpi. Statistical significance was determined by one way ANOVA (n = 14–18, ****p < 0.0001). (F) Infection model for subsequent experiments. |
|
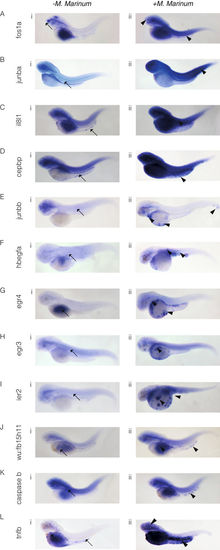
In situ hybridisation reveals spatial analysis of M. marinum-induced upregulated genes. Embryos were injected at the 32-512-cell stage with M. marinum. Expression patterns and gene transcript levels were analysed by in situ hybridisation using digoxigenin-labelled RNA probes, in uninjected controls (A–Li) and M. marinum-injected embryos (A–Lii). Control expression patterns are indicated by the black arrows with black arrowheads used to indicate altered expression patterns for fos1a (A), junba (B), il81a (C), cepbp (D), junbb (E), hbegfa (F), egr4 (G), egr3 (H), ier2 (I), wu:fb15h11 (J), caspase b (K) and tnfb (L). EXPRESSION / LABELING:
PHENOTYPE:
|
|
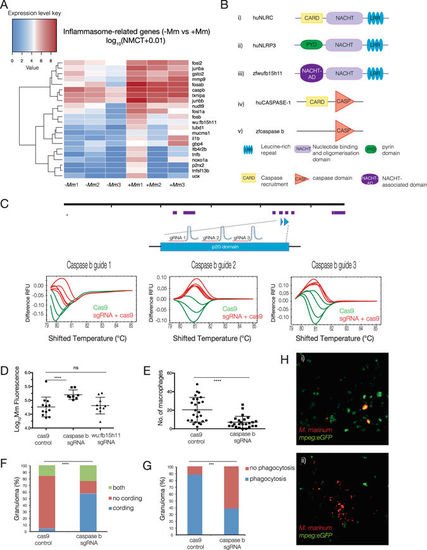
Neutrophils mediate M. marinum killing by inflammasome-dependent mechanism. (A) Heatmap shows the log10 (normalised counts (NMCT) +0.01) of selected differentially expressed transcripts of known inflammasome components and related genes (adjusted p-value < 0.05). Red - high expression. White - medium expression. Blue - low expression. (B) Schematic representation of the domain organisation of huNLRP3 (i), huNLRC (ii), zfwu:fb15h11 (iii), huCaspase 1 (iv) and zfcaspase b (v). (C) caspase b sgRNA tests for efficiency in inducing DNA double-strand breaks. sgRNAs were injected into the 1-cell stage and genomic DNA extracted at 24 hpf. High resolution melting curves from 3–4 individual embryos injected either with Cas9 mRNA only (green) or co-injected with Cas9 mRNA and caspase b sgRNA (red) are shown. (D–H) For quantitative analysis of inflammasome components, embryos were injected at the one-cell stage with Cas9 mRNA or sgRNAs to caspase b or wu:fb15h11 followed by injection of M. marinum at the 32–512 cell stage. (D) Bacterial burden of larvae infected at 3 dpi as measured by Mm fluorescence intensity (red). Statistical significance was determined by two-tailed unpaired students t-test with Welch’s correction. For caspase b sgRNA, ****p < 0.0001. (E) Enumeration of macrophage infiltrating M. marinum at 5 dpi. Statistical significance was determined by two-tailed unpaired students t-test with Welch’s correction (****p < 0.0001). (F) Percentage of larvae with cording phenotype at 5 dpi. Statistical significance was determined by chi-squared test (****p < 0.0001). (G) percentage of larvae where macrophages phagocytise M. marinum at 5 dpi. Statistical significance was determined by Fishers exact test (***p < 0.001). (H) Representative maximum intensity projections of larvae with cording bacteria with decreased phagocytosis in caspase b knockout embryos (i) vs non-cording controls (ii). |

ZFIN is incorporating published figure images and captions as part of an ongoing project. Figures from some publications have not yet been curated, or are not available for display because of copyright restrictions. EXPRESSION / LABELING:
PHENOTYPE:
|