|
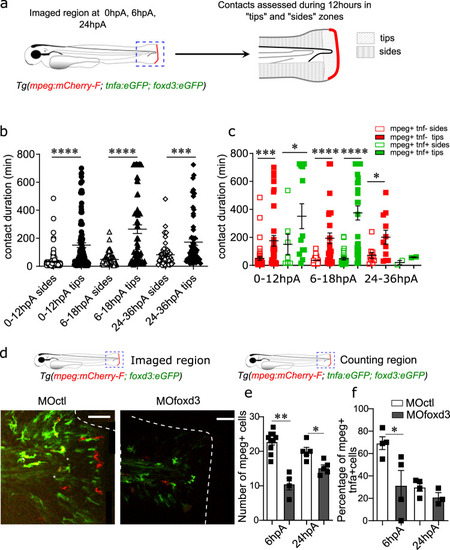
<italic>Foxd3</italic><sup><italic>+</italic></sup> NCdC interact with and promote the recruitment and activation of macrophages in the blastema.a Schematic representation of the tip and sides where macrophages and foxd3+ NCdC contacts were assessed for 12 h by live confocal microscopy at 0hpA, 6 hpA, 24 hpA in Tg(mpeg:mCherry-F; tnfa:eGFP-F; foxd3:eGFP-F) larvae. b Quantification of the duration of macrophages and foxd3+NCdC contacts in the tips and sides during 12 h in Tg(mpeg:mCherry-F; tnfa:eGFP-F; foxd3:eGFP-F) larvae, at 0hpA (n = 192 for the sides group, n = 124 for the tips group), 6 hpA (n = 105 for the sides group, n = 71 for the tips group), and 24 hpA (n = 54 for the sides group, n = 60 for the tips group) (mean number of contact durations ± SEM, from at least n = 5 biologically independent larvae per time points, Kruskal Wallis test with Dunn’s test for multiple comparison were performed, ****p < 0.0001,). c Quantification of the contact duration between macrophages (mpeg + and tnfa+) and foxd3+ cells in tips and sides during 12 h in the transgenic larvae, at 0 hpA, 6 hpA, 24 hpA (mean number of contact durations ± SEM, Kruskal-Wallis test and Dunn’s test for multiple comparison were performed, *p < 0.05, ***p < 0.001, ****p < 0.0001, n = 3 biologically independent larvae per time point). d Fluorescence microscopy images of 3 dpf Tg(mpeg1:mCherry-F/tnfa:eGFP-F/foxd3:eGFP-F) larvae injected with MOctl or MOfoxd3 (Scales bars = 50 µm). e Number of macrophages recruited at the wound site at 6 hpA and 24 hpA in MOctl- and MOfoxd3-injected Tg(mpeg1:mCherry-F) larvae (graph represents mean number of positive cells ± SEM, n = 10 biologically independent larvae for the control group at 6 hpA, n = 4 larvae for the MOfoxd3 group at 6 hpA, n = 5 larvae for both control and morphant group at 24hpA, one-tailed Mann-Whitney test was performed, p = 0.0028 for the 6 hpA time point, p = 0.0106 for the 24 hpA time point, *p < 0.05, **p < 0.01). f Percentage of pro-inflammatory macrophages at the wound site at 6 hpA and 24 hpA (graph represents mean number of mpeg+tnfa+ cells ± SEM, n = 4 biologically independent larvae per groups, one−tailed Mann-Whitney test was performed, p = 0.0286 for the 6 hpA time point, *p < 0.05).
|