- Title
-
CytoCensus, mapping cell identity and division in tissues and organs using machine learning
- Authors
- Hailstone, M., Waithe, D., Samuels, T.J., Yang, L., Costello, I., Arava, Y., Robertson, E., Parton, R.M., Davis, I.
- Source
- Full text @ Elife
|
(A) Diagram of the chamber and sample preparation for long-term time-lapse imaging on an inverted microscope (see Materials and methods). (B) 24 h, confocal 3D time-lapse imaging of a developing larval brain lobe (inset, top left, shows orientation and region of the brain imaged) labelled with Jupiter::GFP and Histone::RFP, and registered over time to account for movement. Arrowheads indicate NBs (magenta) and progeny (cyan), enlarged in the top right insets; a dashed white line indicates the boundary to the optic lobe. (C′) A typical individual dividing NB from a confocal time-lapse image sequence of the brain lobe. The NB is outlined (dashed white line) and indicated with a magenta arrowhead, the progeny (GMC) is indicated by a cyan arrowhead. (C′′) Plot of NB division rate for cultured L3 brains shows that division rate of NBs does not significantly decrease over at least 22 h under imaging conditions (n = 3 brains, not significant (ns), p=0.87, one-way ANOVA), calculated from measured cell cycle lengths. (D′) Typical GMC division in an intact larval brain. The first row of panels shows production of a GMC (cyan arrowhead) by the dividing NB (magenta arrowhead, dashed white outline). Second row of panels, GMCs are displaced over the next 6 to 8 h by subsequent NB divisions, the path of displacement is indicated by the dashed yellow arrow. The last two panels (10 to 18 min) show the division of a GMC (green arrowhead, progeny yellow arrowheads). (D′′) Plot showing the rate of GMC division in the ex vivo brain does not change with time in culture (n = 4 brains, ns, p=0.34, one-way ANOVA), calculated from the number of GMC division events in 4 h. Error bars on plots are standard deviation. Scale bars (B) 50 µm; (C), (D) 10 µm. |
|
Related to |
|
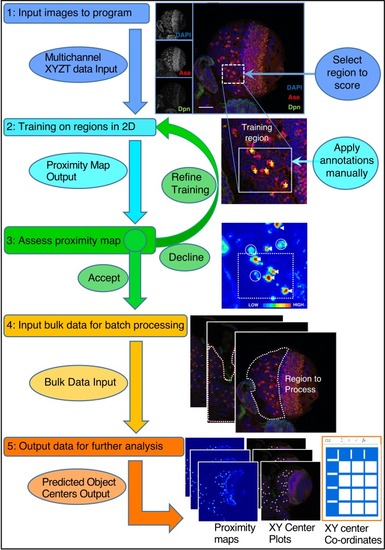
Refer to the Main Text and Materials and methods for details. Training is performed by single click annotation (yellow crosses) within a user-defined region of interest (ROI, white dashed square) to identify the cell class of interest. The resultant proximity map for cell class identification (~probability score for object centres) is evaluated manually to assess the success of training (white arrows indicate good detections and circles indicate where more training may be required). A successful identification regime (Model) is saved and may be used to batch process multiple image data sets. Multiple outputs are produced including a list of the co-ordinates of identified cells. Multiple identification regimes can be sequentially applied to identify multiple cell classes from a single data set. |
|
Related to |
|
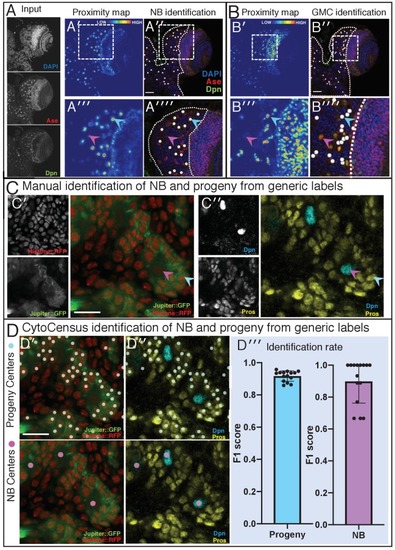
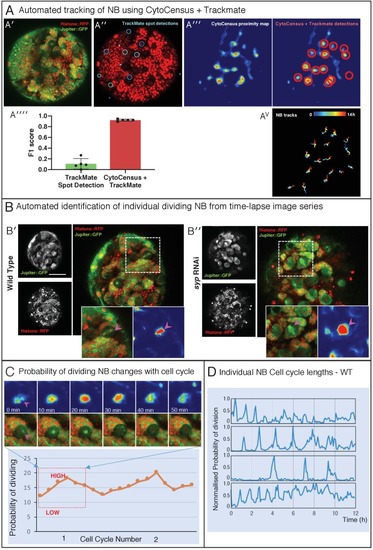
(A) Performance in identifying NBs from 3D confocal image data of a live brain labelled with Jupiter::GFP, Histone::RFP. (A′) Ground Truth manual identification of NB centres. A′′ to ′′′′) Output images comparing NB identification by Ilastik, Fiji-Weka and CytoCensus, white overlay. (Av) Plot comparing object centre detection by TrackMate spot detection, RACE, Fiji-Weka, Ilastik Pixel Classification and CytoCensus (error bars are standard deviation). CytoCensus achieves a significantly better F1-score than Ilastik (p=0.01, n = 3) and FIJI (p=0.005, n = 3). (one-way RM-ANOVA with post hoc t-tests) (B) Comparison of algorithm performance for a 3D neutral challenge data set (B′, see Materials and methods). (B′′, B′′′) Output images comparing object centre determination by Ilastik Pixel Classification and CytoCensus. Segmentation results are shown as green outlines, object centre determination is show as a cyan point. (B′′′′) Plot comparing object centre determination accuracy for the 3D neutral challenge dataset (error bars are standard deviation; p<0.001, Welch’s t-test, n = 25). Scale bars (B) 20 µm; (A′) 50 µm. |
|
Related to |
|
Related to |
|
Related to |
|
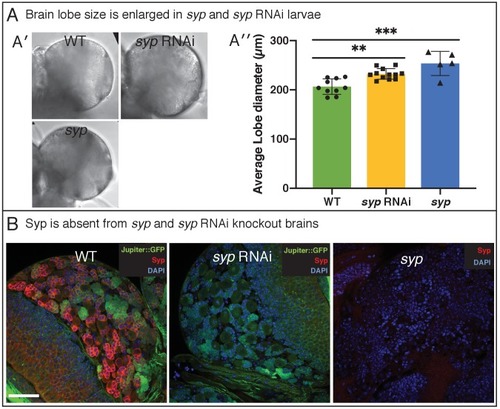
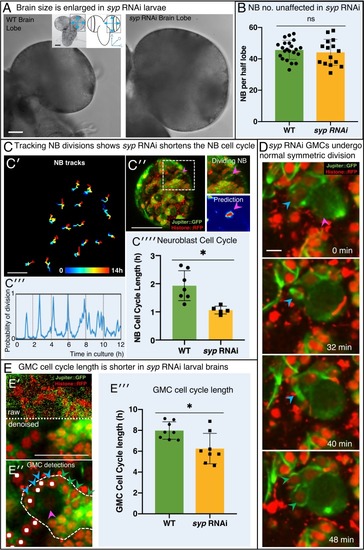
(A) Brightfield images of freshly isolated brains from third instar WT (OregonR) and syp RNAi larvae, respectively. Inserts in (A) show the region of the brain imaged and the measurements taken to compare brain size. (B) Chart comparing NB numbers showing that syp RNAi knockdown does not have a significant effect on NB number/brain (ns, p=0.77, t-test, WT n = 22; RNAi n = 15). NB were identified by Dpn labelling and the average count for a comparable volume of a single optic lobe CB region is shown. (C) Automated identification of NB division using CytoCensus: (C′) Tracking of NB centres, based on CytoCensus detections, over 14 h; (C′′) raw image showing single timepoint from live, 3D time-lapse, confocal imaging (insert = single dividing NB, showing CytoCensus prediction of a dividing NB); (C′′′) graph of division of a single tracked NB over 14 h; (C′′′′) average NB (6–9 NB/brain) cell cycle length is reduced in syp RNAi knockdown brains (p=0.004, Welch’s t-test, WT n = 7, syp RNAi n = 5 brains); (D) Sequence of confocal images from a typical 3D time-lapse movie showing that in syp RNAi brains, GMCs divide normally to produce two equal sized progeny that do not divide further. (E) Semi-automated analysis of GMC division by CytoCensus shows that GMC cell cycle length is reduced in syp RNAi brains. (E′) Single image plane taken from a 3D time-lapse, confocal image data set (imaged at one Z-stack/2 min). showing raw image data (top) and denoised (bottom). (E′′) CytoCensus GMC detections (cyan) with a single NB (magenta), and NB niche (dotted white line), shows GMCs are detected but neurons (green) are not. (E′′′) Plot of GMC cell cycle length, which is decreased in syp RNAi brains compared to WT (p=0.01, Welch’s t-test, n = 8 GMCs from three brains). Scale bars in (A) 50 µm; (C′) 20 µm; (C′′) 50 µm; (D) 5 µm; (E) 25 µm |
|
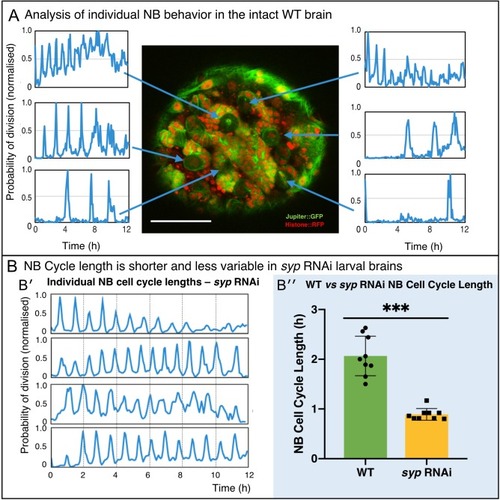
(A) Using the proximity map output of CytoCensus, individual NBs can be followed through their cell cycle. Arrows: Individual NB locations, and the corresponding proximity map output plotted over time for that NB. (B) Comparison of WT and syp RNAi NB: (B′) analysis of cell cycle over time for individual NBs from a syp RNAi brain; (B′′) comparison of cell cycle lengths for individual NB in a single WT vs syp RNAi brain (p=0.002, F-test, n = 9 NB). Scale bar 40 µm |
|
Related to |
|
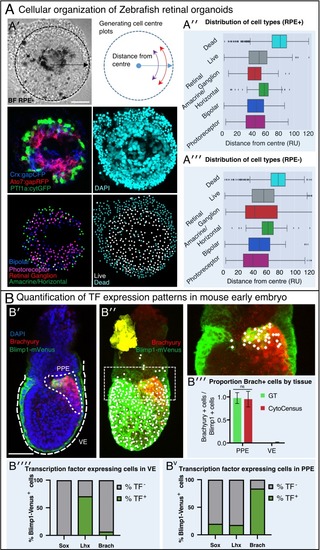
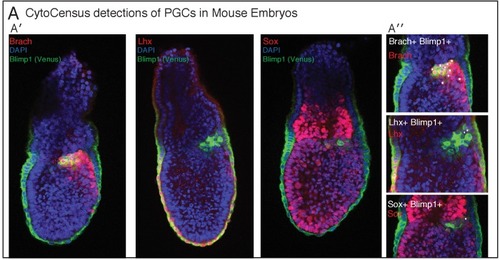
(A) Automated analysis of Zebrafish retinal organoids at the single-cell level. Raw data from Eldred et al. (2017). (A′) Top: brightfield image and diagram indicating the location of cells was defined as displacement from the organoid center. Middle: Cell fate marker expression (Crx:gapCFP; Ato7:gapRFP; PTf1a:cytGFP) and DAPI. Bottom: Cell centre identification by CytoCensus for the different cell types as defined by the labelling profiles (Bipolar, Photoreceptor, Retinal Ganglion, Amacrine/Horizontal, Live/Dead). (A′′-A′′′) Radial distribution of the different cell types determined from cell centre identifications by CytoCensus; the effects on organoid organisation of the presence (A′′) or absence (A′′′) of retinal pigment epithelium (RPE) cells is examined (ns, one-way ANOVA). RU = Radial Units, normalised to a radius of 100 (see Materials and methods) (B) Automated quantification of TF expressing cells in a fixed early streak stage mouse embryo (e6.5) labelled for transcription factors, Blimp1-mVenus and DAPI. (B′) A medial confocal section showing Brachyury in the primitive streak in the proximal posterior epiblast (PPE) and visceral endoderm (VE, highlighted cortical tracing). (B′′) Cortical image of the same mouse embryo overlaid with total cell centre predictions by CytoCensus of Brachyury positive cells; insert to the right is a zoomed in image of the highlighted rectangle showing only cell centre predictions in a single medial plane. (B′′′) Comparison of CytoCensus and manual Ground Truth (GT) measurements of the proportion of Brachyury positive cells from 2D planes in the VE and PPE (ns, t-test, n = 3). (B′′′′-Bv) Proportion of transcription factor positive cells (TF) in, using CytoCensus measurements in 3D according to tissue regions (PPE and VE) defined in (B′). Scale bars 25 µm in (A); 100 µm in (B′). |
|
Related to |