Fig. 3
Fig. 3
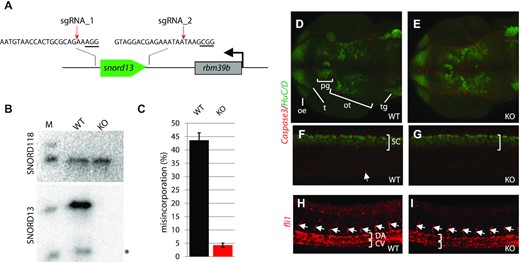
Phenotypic analyses of SNORD13-deficient zebrafish embryos. (A) Schematic representation of the zebrafish snord13 locus. The relative location of the two DNA sequences targeted by sgRNAs is shown (PAM sequence is underlined). (B) Northern blotting showing that SNORD13 is no longer detected in embryos bearing deletion events. *: truncated SNORD13 form which is routinely detected. SNORD118 was used as a gel loading control. (C) Histograms show percentage of misincorporation (C-to-U ratio) measured in borohydride-treated RNA samples prepared from WT and SNORD13-KO embryos (n = 3). (D–G) Confocal projections of WT (D, F) and SNORD13-KO (E, G) embryos after immunostaining against cleaved Caspase-3 (red) and the pan-neural marker HuC/D (green). (D, E) Dorsal view of the zebrafish brain at 30 hpf, neurons from the olfactory epithelium (oe), tectum (t), pineal gland (pg), optic tectum (ot) and trigeminal ganglion (tg) can be detected. (F, G) Lateral view of the spinal cord (sc) at 30 hpf. The white arrow indicates an apoptotic cell stained by activated caspase-3. (H, I) Confocal projections of WT (H) and SNORD13-KO (I) embryos after in situ RNA hybridization using antisense riboprobes directed against fli1 transcripts. Note that in the absence of antibodies to FLI1 that give reliable signals, fli1 mRNA expression was used as a proxy for monitoring vessel development. It does not imply that fli1 expression per se may be altered in SNORD13-KO zebrafish, whether at the mRNA or protein levels. White brackets indicate the dorsal aorta (DA) and the cardinal vein (CV) and the white arrows point to intersegmental vessels. Pictures are representative of three independent experiments (with at least 6 embryos imaged per genotype).