Figure 7
- ID
- ZDB-FIG-210219-22
- Publication
- Prentzell et al., 2021 - G3BPs tether the TSC complex to lysosomes and suppress mTORC1 signaling
- Other Figures
- All Figure Page
- Back to All Figure Page
|
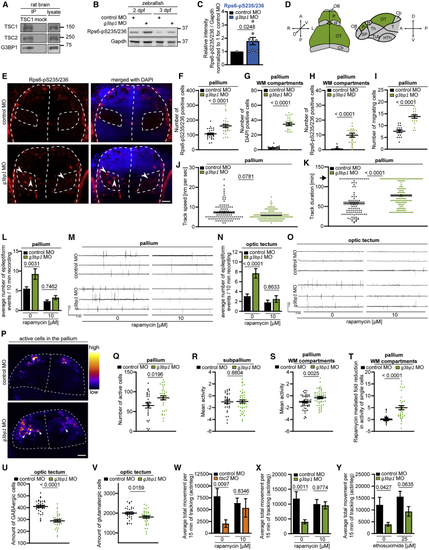
G3BP1 deficiency elicits mTORC1-driven neuronal phenotypes (A) IP against TSC1 (TSC1 #3) or mock (rabbit IgG). n = 2. (B) Zebrafish larvae injected with (C) Quantitation of Rps6-pS235/236 in (B), pooled for 2+3 dpf. Shown are data points and mean ± SEM. (D) Dorsal and lateral view of a zebrafish larva brain. P, pallium; OT, optic tectum; H, habenula; Cb, cerebellum; OB, olfactory bulb; SP, subpallium; Th, thalamus; Tub, tuberculum; T, tegmentum; HTh, hypothalamus. (E) IF of Rps6-pS235/236 in (F) Quantitation of Rps6-pS235/236-positive cells in the pallium in (E). Shown are data points and mean ± SEM. (G) Quantitation of cells in the WM in (E). Data are shown as in (F). (H) Quantitation of Rps6-pS235/236-positive cells in the WM in (E). Data are shown as in (F). (I) Quantitation of HuC-positive cells in (J) Movement speed of single HuC-positive cells. Data are shown as in (I). (K) Track duration of single HuC-positive cells. Data are shown as in (I). Arrow, maximum track duration. (L) Quantitation of epileptiform events in LFP recordings from the pallia of (M) Representative LFP recordings for (L). (N) Quantitation of epileptiform events in LFP recordings from optic tecta of (O) Representative LFP recordings for (N). (P) Neuronal activity in pallia of (Q) Quantitation of active neuronal cells in (P). Shown are data points and mean ± SEM. (R) Quantitation of mean neuronal activity in the subpallia of (S) Quantitation of mean neuronal activity in the WM of (T) Quantitation of rapamycin-mediated fold reduction in the activity of single cells in the WM of (U) Quantitation of GABAergic cells in optic tecta of (V) Quantitation of glutamatergic cells in optic tecta of (W) Locomotor activity of (X) Locomotor activity of (Y) Locomotor activity of See also |
Reprinted from Cell, 184(3), Prentzell, M.T., Rehbein, U., Cadena Sandoval, M., De Meulemeester, A.S., Baumeister, R., Brohée, L., Berdel, B., Bockwoldt, M., Carroll, B., Chowdhury, S.R., von Deimling, A., Demetriades, C., Figlia, G., Genomics England Research Consortium, de Araujo, M.E.G., Heberle, A.M., Heiland, I., Holzwarth, B., Huber, L.A., Jaworski, J., Kedra, M., Kern, K., Kopach, A., Korolchuk, V.I., van 't Land-Kuper, I., Macias, M., Nellist, M., Palm, W., Pusch, S., Ramos Pittol, J.M., Reil, M., Reintjes, A., Reuter, F., Sampson, J.R., Scheldeman, C., Siekierska, A., Stefan, E., Teleman, A.A., Thomas, L.E., Torres-Quesada, O., Trump, S., West, H.D., de Witte, P., Woltering, S., Yordanov, T.E., Zmorzynska, J., Opitz, C.A., Thedieck, K., G3BPs tether the TSC complex to lysosomes and suppress mTORC1 signaling, 655-674.e27, Copyright (2021) with permission from Elsevier. Full text @ Cell