Fig. 4
- ID
- ZDB-FIG-240109-99
- Publication
- Song et al., 2023 - Structural basis for inactivation of PRC2 by G-quadruplex RNA
- Other Figures
- All Figure Page
- Back to All Figure Page
|
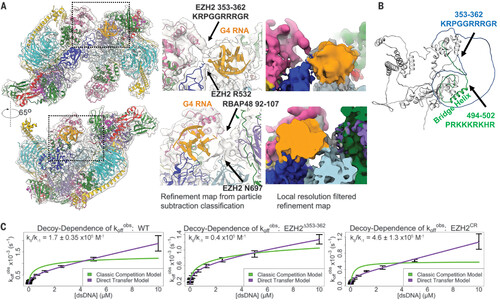
EZH2 loops physically contact G4 RNA and contribute to direct handoff from RNA to DNA. (A) Map of PRC2-1G4 RNA from particle subtraction and classification (fig. S6) with zoom-ins to emphasize the observed physical interactions of PRC2 and G4 RNA. (B) EZH2 structure from AlphaFold predicts two disordered loops of EZH2. Arginine-rich loop [EZH2(353–362)] and lysine-rich loop [EZH2(494–502)] are indicated in blue and green, respectively. (C) FP assays to monitor the transfer kinetics of PRC2 from fluorescently labeled 1G4 RNA to a dsDNA competitor. The ratio kθ/k–1 of the PRC2 RNA-to-dsDNA direct-transfer rate constant (kθ) and the PRC2-RNA dissociation rate constant (k–1) provides a measure of the propensity of PRC2 to exchange these ligands by the direct transfer mechanism. WT PRC2 had kθ = 90 ± 11 M−1s−1, k–1 = 5.6 ± 0.49 × 104 s−1, and kθ/k–1 = 1.7 ± 0.35 × 105 M−1. EZH2 ∆353–362 had kθ = 48 M−1s−1, k–1 = 12 × 104 s−1, and kθ/k–1 = 0.4 × 105 M−1. EZH2 CR had kθ = 130 ± 62 M−1s−1, k–1 = 2.7 ± 0.63 × 104 s−1, and kθ/k–1 = 4.6 ± 1.3 × 105 M−1. koffobs, dissociation rate constant observed. |