Fig. S1
|
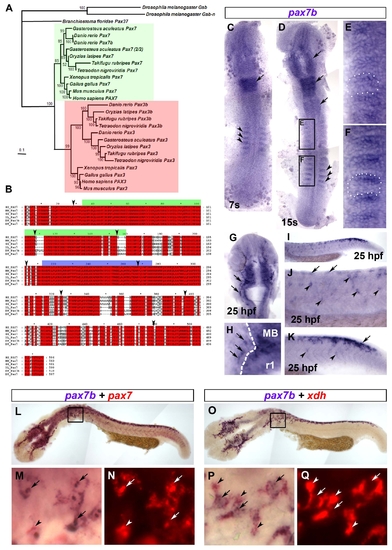
Phylogenetic analysis and expression of pax7b. A. Bayesian phylogenetic analysis of the Pax3/7 sub-family. The tree was rooted with D. melanogaster Gsb and Gsb-n. Confidence for nodes is indicated by Bayesian support values. Green and red boxes indicate Pax7 and Pax3 sub-groups, respectively. B. Multiple sequence alignment of Pax7b sequence with homologous Pax7 proteins. The green and blue bars above the sequences indicate the Paired box and homeodomain, respectively. Arrowheads denote exon boundaries. Red boxes denote identical residues. Grey boxes show similar residues. HS = Homo sapiens, MM = Mus musculus, GG = Gallus gallus, XL = Xenopus leavis, DR = Danio rerio. C-Q. In situ mRNA hybridisation analysis of pax7b (C-Q) alone or co-localised with either pax7 (L-N) or xdh (O-Q), Dorsal view flatmounts are anterior to top (C-H) and lateral view flatmounts are anterior to left and dorsal to top (I-K,L-Q). At 7s, pax7b is expressed in anterior rhombomeres (C,D arrow) and the anterior edge of the newly-formed somites (C,D, arrowheads). By 15s, pax7b expression is maintained in rhombomeric stripes and appears in midbrain and anterior spinal cord (D, arrows). Expression in somites (black boxes, magnified in E,F, white dots delineate a somite) is reduced in older somites (E), but abundant in the anterior region of young somites (F). At 25 hpf, pax7b is expressed in cranial NC (G,H arrows; MB = midbrain, r1 = rhombomere 1; white dashes CNS border). In trunk NC, pax7b is expressed primarily in nascent pre-migratory cells (arrows) in tail (I enlarged in K) and older migratory cells (arrowheads) in trunk (I enlarged in J). Pax7b co-localises with both pax7 (L-N) and xdh (O-Q) mRNA. Boxes (L,O) are magnified in M,N,P,Q to show regions of overlapping (arrows), and non-overlapping (arrowheads) mRNA. Xdh mRNA is mainly cytoplasmic (arrowheads, P,Q), whereas pax7b and pax7 mRNAs show prominent nuclear puncta (arrows, M,P). |
| Genes: | |
|---|---|
| Fish: | |
| Anatomical Terms: | |
| Stage Range: | 5-9 somites to Prim-5 |
Reprinted from Developmental Biology, 317(2), Minchin, J.E., and Hughes, S.M., Sequential actions of Pax3 and Pax7 drive xanthophore development in zebrafish neural crest, 508-522, Copyright (2008) with permission from Elsevier. Full text @ Dev. Biol.