- Title
-
siRNAs Targeting Non-Human Species-Specific lncRNAs Trigger Cell Death in Human Colorectal Cancer Cells
- Authors
- Feng, W.Y., Zeng, J.X., Chen, Y.R., Fang, Z.P., Gao, Y., Zhou, W.J.
- Source
- Full text @ J Cancer
|
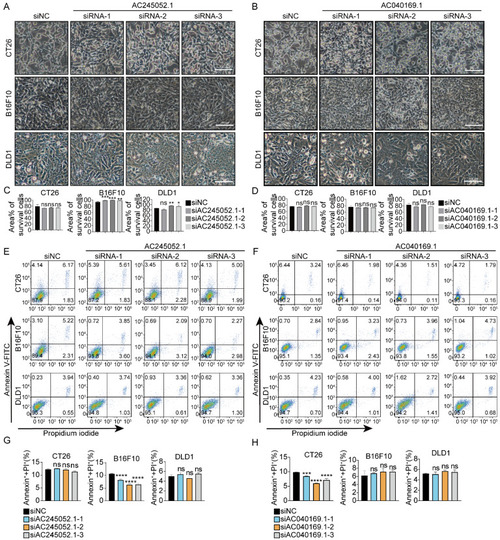
siRNAs designed to target mouse-specific lncRNAs lead to cell death in human CRC cells ( |
|
Targeting human-specific lncRNAs with siRNAs fails to trigger cell death in mouse cancer cells ( |
|
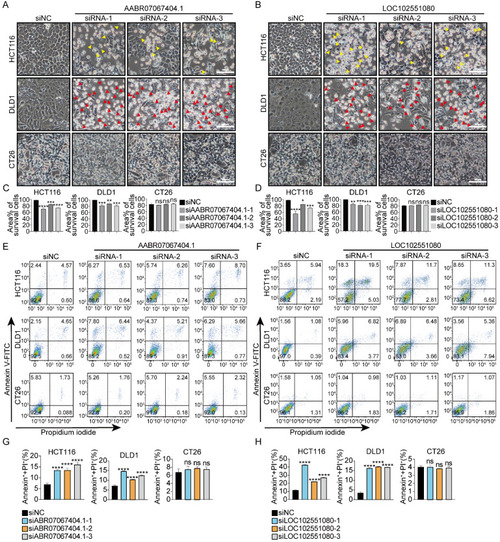
siRNAs targeting rat-specific lncRNAs exhibit specific cytotoxicity against human cancer cells without affecting mouse cancer cells ( |
|
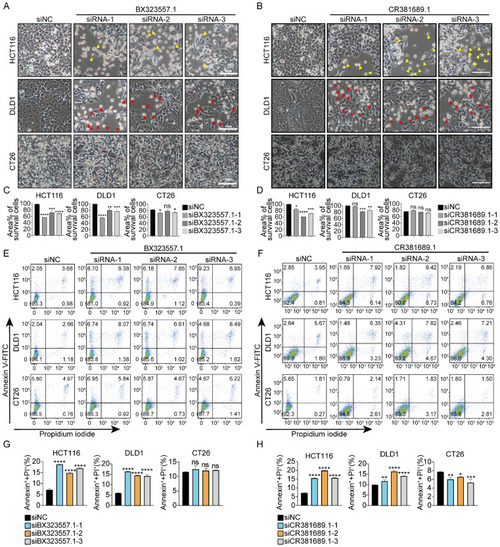
siRNAs targeting zebrafish-specific lncRNAs exhibit specific cytotoxicity against human cancer cells without affecting mouse cancer cells ( |
|
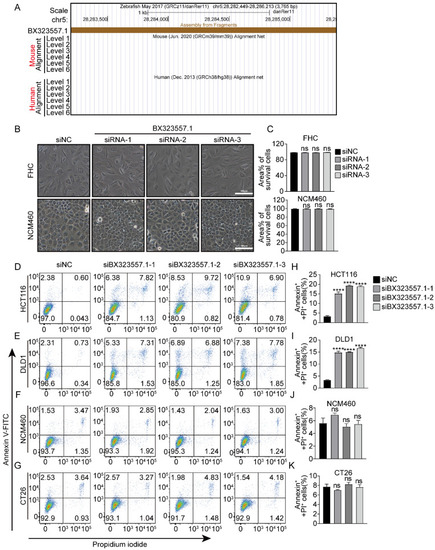
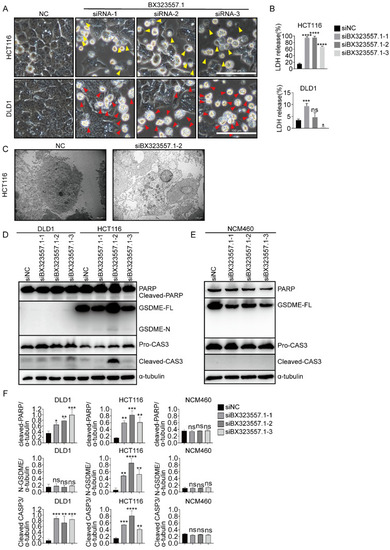
siRNAs targeting zebrafish-specific lncRNA BX323557.1 do not cause death in normal human or mouse cancer cells ( |
|
siRNAs designed to target zebrafish-specific lncRNA BX323557.1 prompt apoptosis or pyroptosis in human CRC cells ( |
|
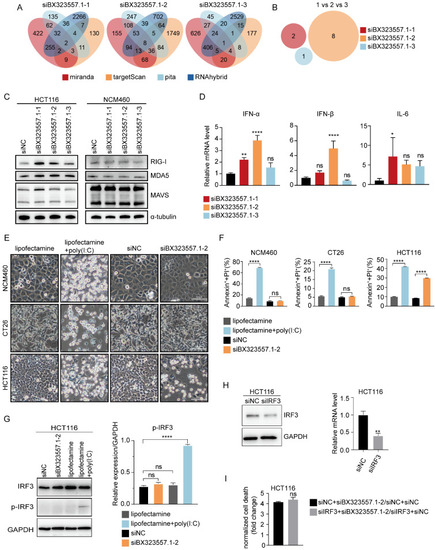
siRNAs targeting zebrafish-specific lncRNA BX323557.1 elicit cell death by triggering an IRF3-independent immune response. ( |