- Title
-
Zebrafish Retinal Ganglion Cells Asymmetrically Encode Spectral and Temporal Information across Visual Space
- Authors
- Zhou, M., Bear, J., Roberts, P.A., Janiak, F.K., Semmelhack, J., Yoshimatsu, T., Baden, T.
- Source
- Full text @ Curr. Biol.
|
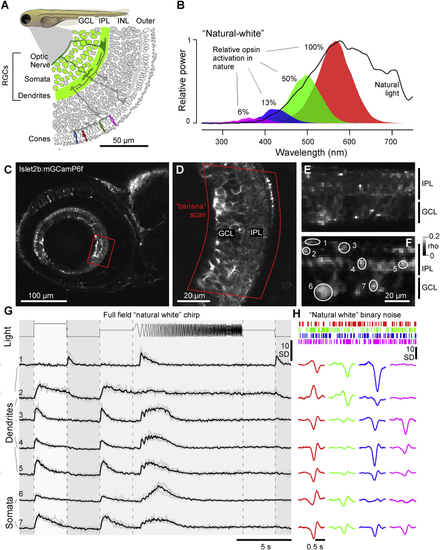
Recording from RGC Dendrites and Somata (A) Schematic of Islet2b:mGCaMP6f expression in RGCs (green) across a section of the larval zebrafish eye, with somata in the ganglion cell layer (GCL) and dendrites in the inner plexiform layer (IPL); see also (B) Average spectrum of natural daylight measured in the zebrafish natural habitat from the fish’s point of view along the underwater horizon (solid line). Convolution of the zebrafish’s four cone action spectra with this average spectrum (shadings) was used to estimate the relative power each cone surveys in nature, normalized to red cones (100%). Stimulation LED powers were relatively adjusted accordingly (“natural white”). (C and D) GCaMP6f expression under two-photon surveyed across the entire eye’s sagittal plane (C) and zoom-in to the strike zone as indicated (D). Within the zoomed field of view, a curved scan path was defined (“banana scan”) to follow the curved GCL and IPL for activity recordings (E), which effectively “straightened” the natural curvature of the eye. (E and F) Example activity scan with RGC dendrites occupying the top part of the scan in the IPL and somata occupying the bottom part in the GCL as indicated (E) and correlation projection [ (G) Mean (black) and individual repeats (gray) example responses of ROIs from (E) to full-field stimulation as indicated. (H) As (G), now showing linear kernels to red, green, blue, and UV components recovered from natural white noise stimulation ( Note that several ROIs display a robust UV component despite the ~20-fold attenuated stimulation power in this band relative to red (B). See also |
|
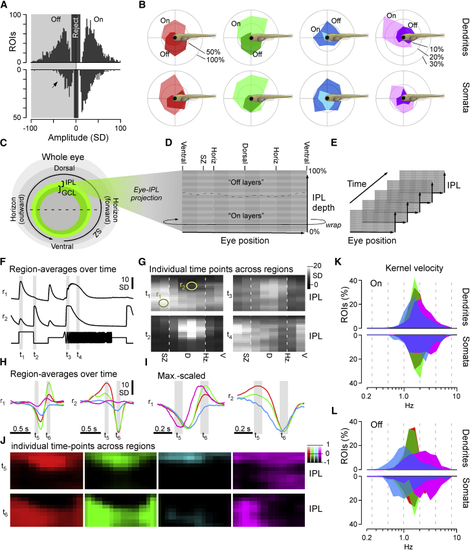
Major Functional Response Trends across the Eye (A) Kernel amplitudes of all dendritic (top) and somatic (bottom; y-flipped) ROIs, shown for the maximal amplitude kernel of each ROI irrespective of color. For a breakdown by color, see (B) Prominence of different color and polarity responses among dendrites (top row) and somata (bottom row), plotted across visual space. In each case, all kernels that exceeded a minimum amplitude of 10 SDs were included. Scale bars in percent of dendritic/somatic ROIs that were recorded in a given section of the eye such that the percentages of On, Off, and non-responding (<10 SD) add to 100% are shown. (C–E) Schematic illustrating how dendritic ROIs from different parts of the eye and IPL depth (C) were mapped into a 2D “Eye-IPL” map (D), which can then also be analyzed over time (E). Note that this involved “cutting” the circular range of eye positions such that the ventral retina is represented at either edge along the 2-projections’ x axis. (F and G) Example snapshots of mean responses to chirp stimulation (cf. (H–J) As (F) and (G) but instead showing mean kernels across the four spectral wavebands, where (H) and (I) are mean and max-scaled mean kernels for Eye-IPL regions r1,2 (as in F), respectively. (J) shows each kernel’s full Eye-IPL map at two time points t5,6 as indicated in (H) and (I) (see also (K and L) Distribution of central frequencies ( |
|
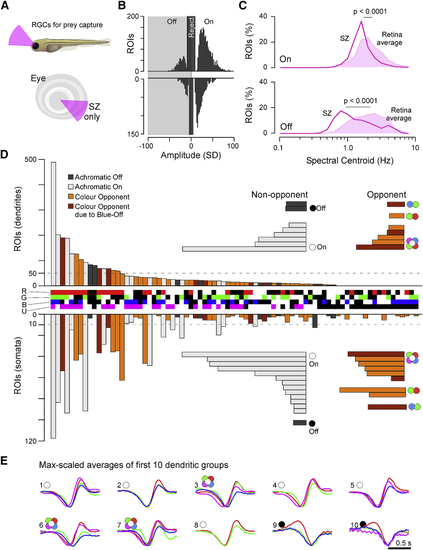
Diverse Color Opponencies in RGCs (A) Each dendritic (top) and somatic (bottom; inverted y axis) ROI that passed a minimum response criterion ( (B) Maximum-amplitude scaled average kernels of the ten most abundant spectral classes among dendrites in (A). (C and D) Dendritic groups from (A) summarized according to their position in an Eye-IPL map (cf. |
|
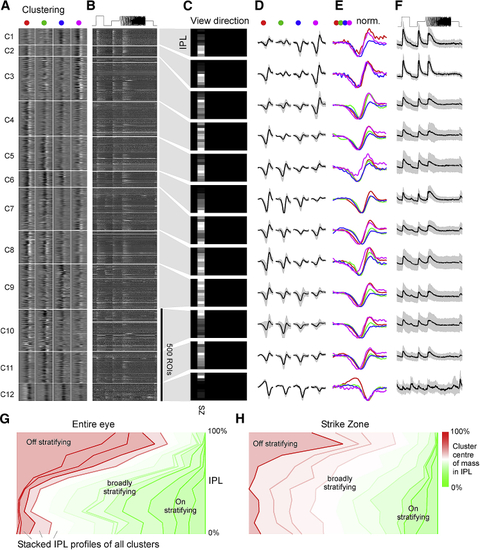
Functional Clustering of Dendritic ROIs (A–F) Dendritic ROIs from across the entire eye were clustered based on their four spectral kernels ( (G) Summary of cluster distributions across the eye, irrespective of IPL depths, for dendritic (top) and somatic (bottom) clusters, scaled by their relative abundance (in %; see scale bars). Eye-distribution profiles were manually allocated to one of the following groups based on which part of visual space is mainly surveyed: SZ (dendritic C1; somatic C2); forward (dendritic C5; somatic C3); outward (dendritic C3,9; somatic C9,11); horizon (dendritic C2,11; somatic C1,4,10); up (dendritic C4–8; somatic C7); and down (dendritic C10,12–15; somatic C12,13). Two large clusters (somatic C5,8) did not obviously fit to any of these categories and were instead grouped separately as “mixed.” It is possible that these clusters comprise several smaller groups of functional RGCs with distinct eye-wide distributions. (H) As (E) for both dendritic (top) and somatic (bottom) data, but with all spectral kernels in each waveband superimposed. Note kinetic similarities across most red and green kernels and near complete absence of positive deflections in blue kernels. |
|
RGC Circuits in the Strike Zone (A) A second series of RGC imaging experiments as shown in (B) Overview of dominant On and Off responses among dendrites (top) and somata (bottom) for the SZ. Dendrites n = 2,370 On, n = 624 Off; somata n = 1,312 On, n = 379 Off. Chi-square with Yates correction for On:Off distributions dendrites versus somata: p < 0.22. For details, cf. (C) Relatively slowed central frequency tuning of SZ-UV kernels (lines) compared to the retina average of UV kernels (filled) among both On (top) and Off (bottom) kernels (cf. (D) Ternary spectral classification of SZ dataset (for details, cf. (E) Maximum amplitude scaled average kernels of the ten most abundant spectral classes among dendrites in (D). |
|
The SZ Is Dominated by Broadly Stratifying UV-Sensitive On Clusters (A–F) Clustering of dendritic ROIs from SZ dataset (for details, cf. (G and H) Side-to-side comparison of functional stratification profiles of clusters from data across the eye (G; cf. |
|
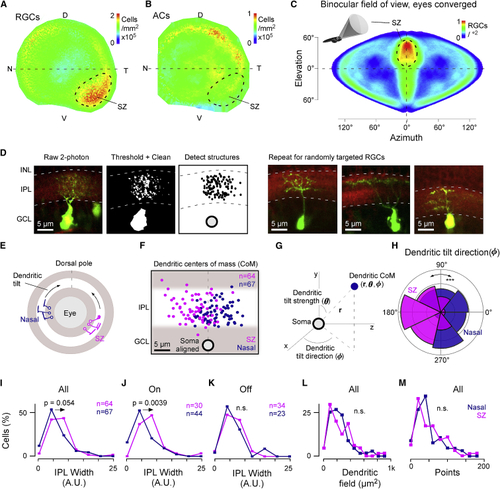
Elevated RGC Density and Relative Overrepresentation of Diffuse ON-RGCs in the SZ (A and B) Density maps of all RGCs (A) and ACs (B) computed from cell counts in (C) Projections of RGC densities from (A) into binocular visual space during hunting (eyes converged), as illustrated in the inset. Note that the two SZs neatly superimpose (see also [ (D) Illustration of photoconversion and pre-processing pipeline for digitizing single RGC morphologies. Left: following photoconversion, cells were imaged as stacks under two-photon (green) in the background of BODIPY staining to demarcate the IPL borders (red). Cells were then thresholded and manually “cleaned” where required prior to automatic detection of image structures and alignment relative to the IPL borders. The resultant “point clouds” were used to determine summary statistics of each cell (e.g., E–M) and were also projected into density maps for visualization ( (E–M) A total of n = 64 and n = 67 randomly targeted RGCs from the SZ and nasal retina, respectively, were processed for further analysis, which included computation of their dendritic tilt (E–H), stratification widths within the IPL (I–K), |