- Title
-
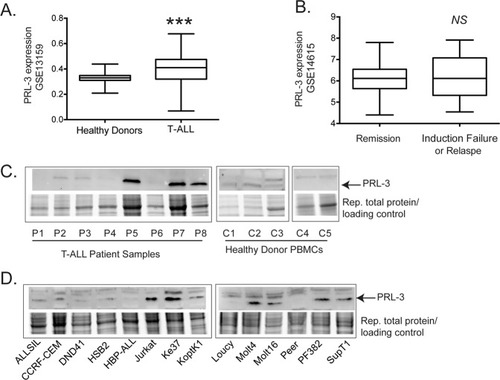
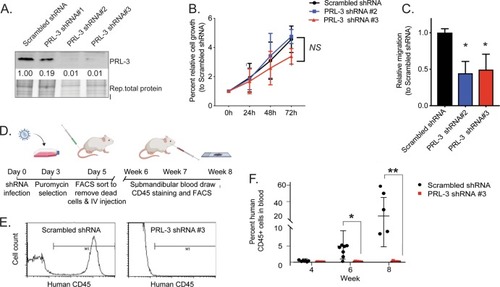
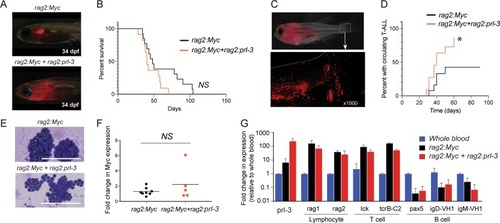
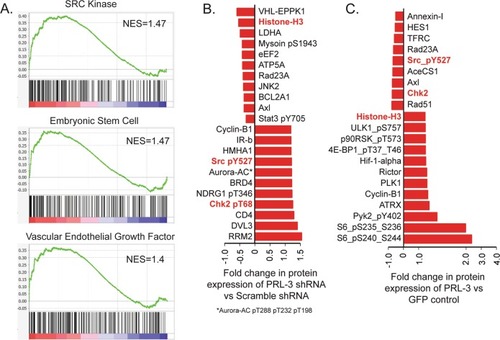
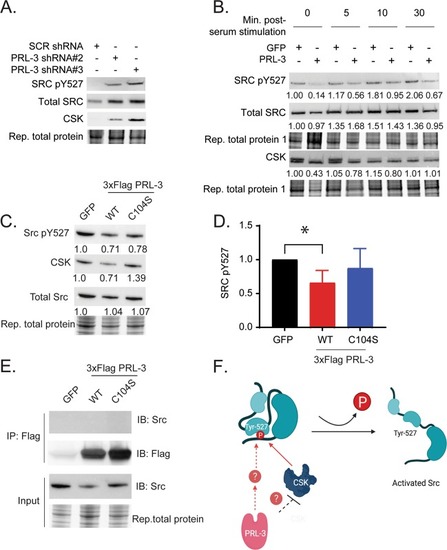
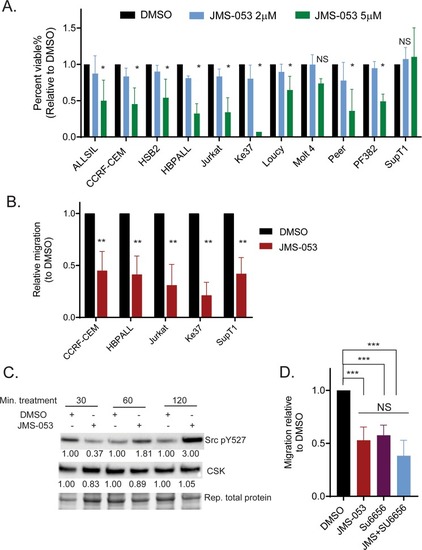
Protein tyrosine phosphatase 4A3 (PTP4A3/PRL-3) drives migration and progression of T-cell acute lymphoblastic leukemia in vitro and in vivo
- Authors
- Wei, M., Haney, M.G., Rivas, D.R., Blackburn, J.S.
- Source
- Full text @ Oncogenesis
|
|
|
|
|
|
|
|
|
|
|
|