Fig. 1
- ID
- ZDB-FIG-250807-26
- Publication
- Wang et al., 2025 - EPHA4 signaling dysregulation links abnormal locomotion and the development of idiopathic scoliosis
- Other Figures
- All Figure Page
- Back to All Figure Page
|
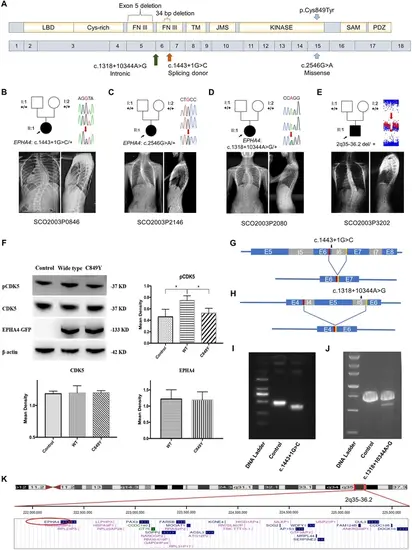
Clinical and genetic information on idiopathic scoliosis (IS) patients and functional effect of EPHA4 variants. (A) Locations of three EPHA4 single-nucleotide variants relative to the protein domains (top panel) and exons 1–18 (bottom panel). (B-E) Pedigrees and spinal radiographs of four probands with dominant gene variants. Sanger sequencing confirmed the variants. The arrows indicate the probands. The term +/+ denotes the wild-type, and cDNA change/+ denotes the heterozygous variant. (F) Western blot analysis of EPHA4-c.2546G>A variant showing the protein expression levels of EPHA4 and CDK5 and the amount of phosphorylated CDK5 (pCDK5) in HEK293T cells transfected with EPHA4-mutant or EPHA4-WT plasmid. WT: wild-type. Data represent three independent experiments. Error bars show mean ± SD. P<0.05 was considered statistically significant. (G) Schematic representation of the effect of the EPHA4-c.1443+1G>C mutation on the splicing process. This variant induced a new splicing site (red box). The yellow box indicates the splicing acceptor. (H) Schematic representation of the effect of the EPHA4-c.1318+10344A>G mutation on the splicing process. This variant induced a new splicing site (red box). The yellow box indicates the splicing acceptor. (I) The minigene assay result showed that the c.1443+1G>C variant introduced a new splicing site, resulting in a 36 bp in-frame deletion in exon 6. (J) The nested PCR showed that the c.1318+10344A>G variant induced exon 5 skipping, resulting in a 339 bp in-frame deletion. (K) NCBI RefSeq genes included in 2q35-q36.2 from UCSC Genome Browser. EPHA4 is shown by the red oval. |