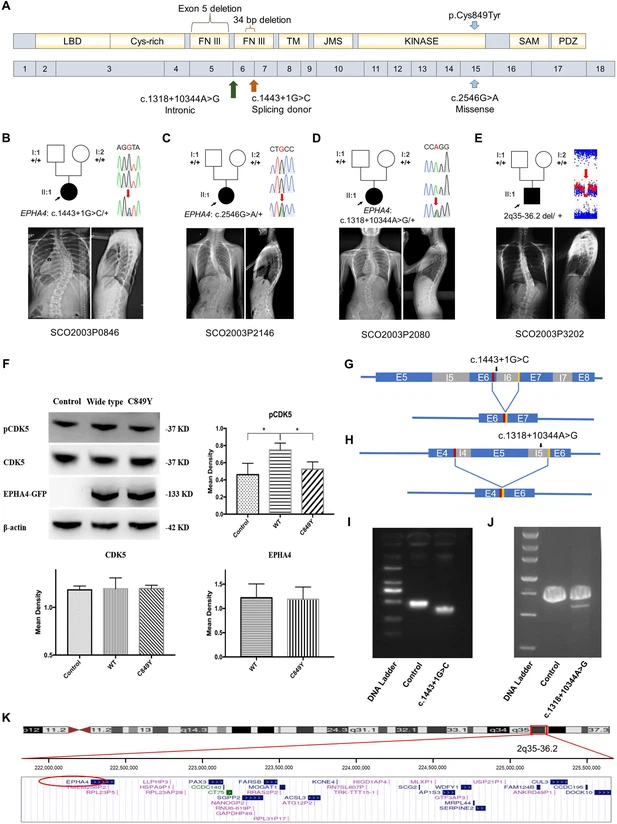
Fig. 1 Clinical and genetic information on idiopathic scoliosis (IS) patients and functional effect of EPHA4 variants. (A) Locations of three EPHA4 single-nucleotide variants relative to the protein domains (top panel) and exons 1–18 (bottom panel). (B-E) Pedigrees and spinal radiographs of four probands with dominant gene variants. Sanger sequencing confirmed the variants. The arrows indicate the probands. The term +/+ denotes the wild-type, and cDNA change/+ denotes the heterozygous variant. (F) Western blot analysis of EPHA4-c.2546G>A variant showing the protein expression levels of EPHA4 and CDK5 and the amount of phosphorylated CDK5 (pCDK5) in HEK293T cells transfected with EPHA4-mutant or EPHA4-WT plasmid. WT: wild-type. Data represent three independent experiments. Error bars show mean ± SD. P<0.05 was considered statistically significant. (G) Schematic representation of the effect of the EPHA4-c.1443+1G>C mutation on the splicing process. This variant induced a new splicing site (red box). The yellow box indicates the splicing acceptor. (H) Schematic representation of the effect of the EPHA4-c.1318+10344A>G mutation on the splicing process. This variant induced a new splicing site (red box). The yellow box indicates the splicing acceptor. (I) The minigene assay result showed that the c.1443+1G>C variant introduced a new splicing site, resulting in a 36 bp in-frame deletion in exon 6. (J) The nested PCR showed that the c.1318+10344A>G variant induced exon 5 skipping, resulting in a 339 bp in-frame deletion. (K) NCBI RefSeq genes included in 2q35-q36.2 from UCSC Genome Browser. EPHA4 is shown by the red oval.
Image
Figure Caption
Acknowledgments
This image is the copyrighted work of the attributed author or publisher, and
ZFIN has permission only to display this image to its users.
Additional permissions should be obtained from the applicable author or publisher of the image.
Full text @ Elife