- Title
-
Biallelic variants in DNAJC7 cause familial amyotrophic lateral sclerosis with the TDP-43 pathology
- Authors
- Yamashita, T., Yokota, O., Ousaka, D., Sun, H., Haraguchi, T., Ota-Elliott, R.S., Matsuoka, C., Kawano, T., Nakashima-Yasuda, H., Fukui, Y., Nakano, Y., Morihara, R., Hasegawa, M., Hosono, Y., Terada, S., Takaki, M., Ishiura, H.
- Source
- Full text @ Acta Neuropathol.
|
Identification of homozygous |
|
Phosphorylated TDP-43 pathology, Bunina bodies, and severe loss of the upper and lower motor neurons in the older brother (IV-1) with a homozygous c.518dupC frameshift variant in |
|
Reduced expression of DNAJC7 protein in the motor cortex of FALS cases with the |
|
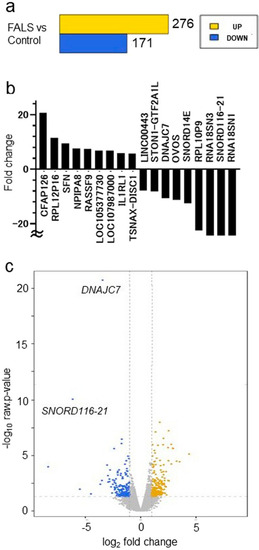
RNA-seq analysis revealed that |
|
|
|
Knockdown of PHENOTYPE:
|