- Title
-
Identification of downstream effectors of retinoic acid specifying the zebrafish pancreas by integrative genomics
- Authors
- López-Pérez, A.R., Balwierz, P.J., Lenhard, B., Muller, F., Wardle, F.C., Manfroid, I., Voz, M.L., Peers, B.
- Source
- Full text @ Sci. Rep.
|
Effect of RA and BMS493 on the transcriptome of endodermal cells. ( |
|
Identification of RAR binding sites in the zebrafish endoderm. ( |
|
Integrated analysis of the ChIP-seq and RNA-seq data. ( |
|
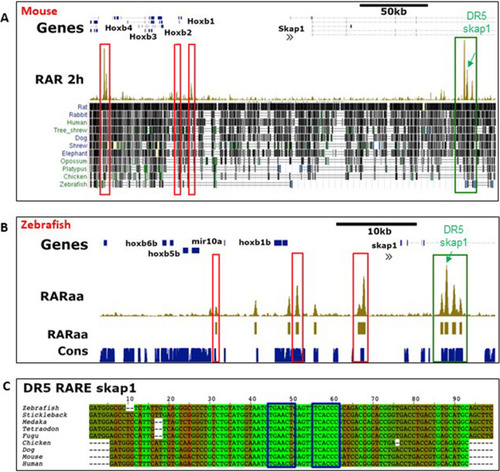
Examples of RARa binding sites conserved among vertebrates. ( |
|
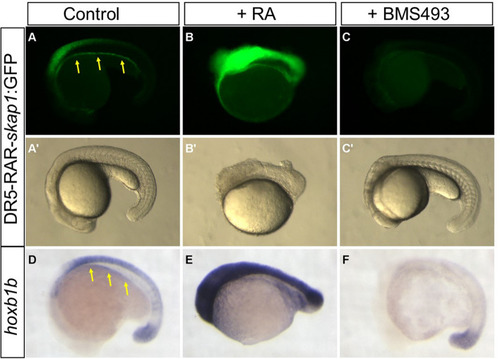
The conserved RAR site from skap1 gene 4th intron is a functional RARE. Pictures of the DR5-RAR-skap1:GFP transgenic embryos treated with DMSO (control, panel |
|
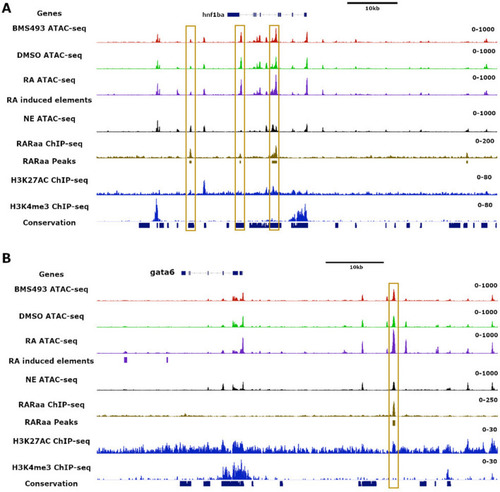
Identification of nucleosome-free regions in zebrafish endodermal cells and following RA treatments by ATAC-seq assays. ( |
|
Location of RARa binding sites and of nucleosome-free regions in the |