- Title
-
Alteration and the Function of Intestinal Microbiota in High-Fat-Diet- or Genetics-Induced Lipid Accumulation
- Authors
- Qiao, F., Tan, F., Li, L.Y., Lv, H.B., Chen, L., Du, Z.Y., Zhang, M.L.
- Source
- Full text @ Front Microbiol
|
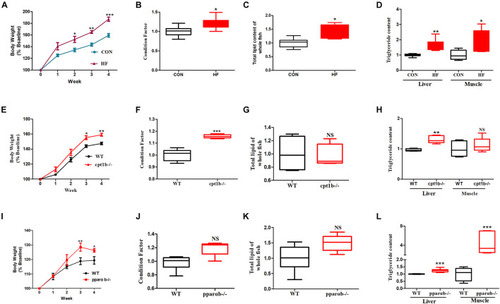
Phenotypes of zebrafish with different genotypes or fed on different diets. (A) Growth curve established using average bodyweight of two dietary groups (CON and HF) every week for 4 weeks (n = 3 per group). (B) Condition factor (CF) of two dietary groups (n = 15 per group). (C) Total lipid of whole fish of two dietary groups (n = 15 per group). (D) Triglyceride contents of liver and muscle of two groups (n = 8 per group). (E) Growth curve established using average bodyweight of WT and cpt1b–/– zebrafish every week for 4 weeks (n = 3 per group). (F) Condition factor of WT and cpt1b–/– zebrafish (n = 15 per group). (G) Total lipid of whole fish of WT and cpt1b–/– zebrafish (n = 15 per group). (H) Triglyceride contents in liver and muscle of WT and cpt1b–/– zebrafish (n = 8 per group). (I) Growth curve established using average bodyweight of WT and pparab–/– zebrafish every week for 4 weeks (n = 3 per group). (J) Condition factor of WT and pparab–/– zebrafish (n = 15 per group). (K) Total lipid of whole fish of WT and pparab–/– zebrafish (n = 15 per group). (L) Triglyceride contents in liver and muscle of WT and pparab–/– zebrafish (n = 8 per group). All data are normalized to control group (100%) and presented as means ± SEM. Significance difference was analyzed by Student’s t-test (*p < 0.05, **p < 0.01, ***p < 0.001). |
|
Boxplots of α diversity indices of bacterial community based on 16S rRNA sequencing. (A–D) Gut microbial α diversity of dietary groups based on (A) ACE, (B) Chao1, (C) Shannon, and (D) Simpson. (E–H) Gut microbial α diversity of WT and cpt1b–/– zebrafish groups based on (E) ACE, (F) Chao1, (G) Shannon, and (H) Simpson. (I–L) Gut microbial α diversity of WT and pparab–/– zebrafish based on (I) ACE, (J) Chao1, (K) Shannon, and (L) Simpson. All data are normalized to control group (100%) (**p < 0.01). |
|
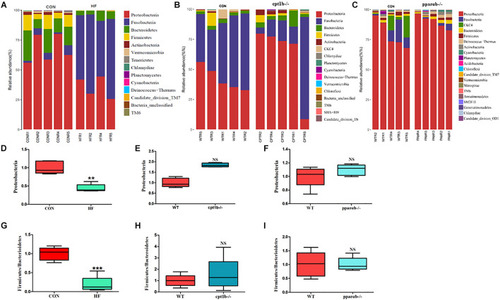
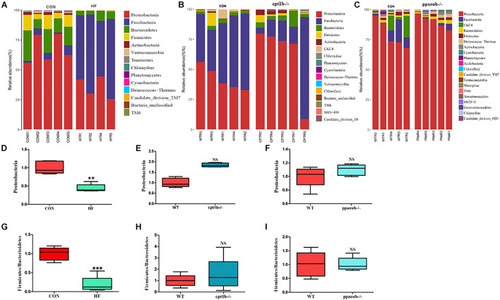
Relative phyla abundance of microbiome based on 16S rRNA gene sequencing. (A) Relative abundances of phylum level among dietary groups (CON and HF). (B) Relative abundances of phylum level between WT and cpt1b–/– zebrafish. (C) Relative abundances of phylum level between WT and pparab–/– zebrafish. (D–F) Ratio of Firmicutes/Bacteroidetes in gut microbiota of (D) dietary groups, (E) WT and cpt1b–/– zebrafish, and (F) WT and pparab–/– zebrafish. (G–I) Ratio of Fusobacteria/Proteobacteria in gut microbiota of (G) dietary groups, (H) WT and cpt1b–/– zebrafish, and (I) WT and pparab–/– zebrafish. All data are normalized to control group (100%) and presented as means ± SEM (**p < 0.01, ***p < 0.001). |
|
Relative phyla abundances of microbiome based on 16S rRNA sequencing. (A) Relative abundances of phylum level of two dietary groups (CON and HF). (B) Relative abundances of phylum level between WT and cpt1b–/– zebrafish. (C) Relative abundances of phylum level between WT and pparab–/– zebrafish. (D–F) Ratio of Firmicutes/Bacteroidetes in gut microbiota of (D) dietary groups, (E) WT and cpt1b–/– zebrafish, and (F) WT and pparab–/– zebrafish. (G–I) Ratio of Fusobacteria/Proteobacteria in gut microbiota of (G) dietary groups, (H) WT and cpt1b–/– zebrafish, and (I) WT and pparab–/– zebrafish. All data are normalized to control group (100%) and presented as means ± SEM (**p < 0.01, ***p < 0.001). |
|
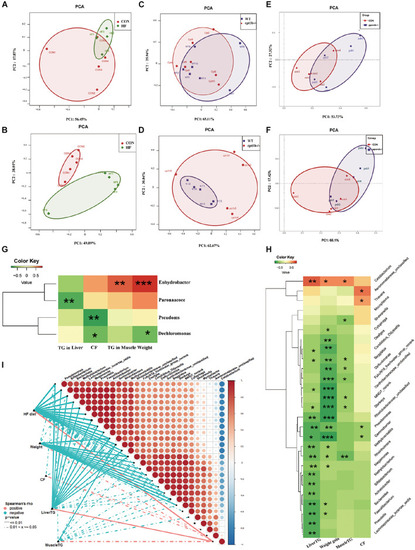
Comparison of gut microbiota composition among different dietary groups and different genotypes on same diets. (A) Principal components analysis (PCA) scores plot of dietary groups (CON and HF) based on 16S rRNA genes (n = 5 per group). (B) PCA scores plot of dietary groups (CON and HF) based on 16S rRNA (n = 5 per group). (C) PCA scores plot of WT and cpt1b–/– zebrafish based on 16S rRNA genes (n = 5 per group). (D) PCA scores plot of WT and cpt1b–/– zebrafish based on 16S rRNA (n = 5 per group). (E) PCA scores plot of WT and pparab–/– zebrafish based on 16S rRNA genes (n = 5 per group). (F) PCA scores plot of WT and pparab–/– zebrafish based on 16S rRNA (n = 5 per group). (G,H) Heatmap illustrates Spearman’s correlation between microbial genus and lipid overaccumulation traits measured between control and high-fat diet group based on (G) total microbial analysis and (H) active microbial analysis. (I) Correlation between abundance of microbial genus and lipid overaccumulation traits in HF groups based on active microbial analysis. *p < 0.05, **p < 0.01, ***p < 0.001. |
|
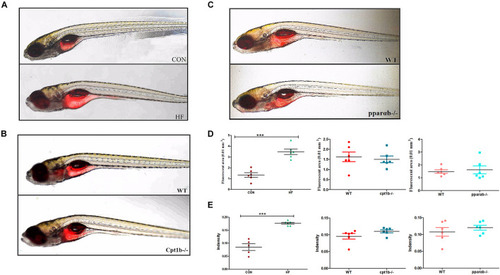
Nile red staining visualizes deep tissue fat deposition in zebrafish. (A–C) Fluorescent images of adipose tissue using Nile red staining of (A) CON and HF zebrafish, (B) WT and cpt1b–/– zebrafish, and (C) WT and pparab–/– zebrafish. (D,E) Quantitative evaluation of fluorescent area (D) and intensity (E) of Nile red staining (n = 6 per group). All data are presented as means ± SEM (***p < 0.001). |