Fig. 7
- ID
- ZDB-FIG-240506-44
- Publication
- Jia et al., 2024 - Metabolome evidence of CKDu risks after chronic exposure to simulated Sri Lanka drinking water in zebrafish
- Other Figures
- All Figure Page
- Back to All Figure Page
|
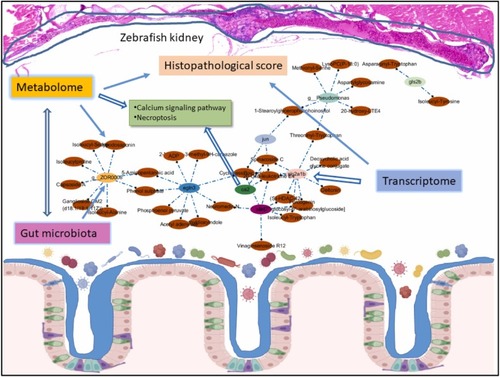
The potential mechanisms of environmental factor-induced CKDu risks based on zebrafish omics changes. Based on the correlation analyses of metabolites and environmental factors, H&E score, DEGs, and gut bacteria at phylum and genus level, the network for multiple relationships was detected and screened with Spearman analysis of the total different metabolites set. Then, the images of the network were constructed by using the String website and Cytoscape (Version 3.10.0). The omics analyses discovered that the gut bacteria of g_ZOR0006, g_Pseudomonas, and DEGs of egln3, ca2, jun, slc2a1b, gls2b, etc, were markedly closely related to metabolites of 20-hydroxy-LTE4, PS(18:0/22:2(13Z,16Z)), neuromedin N, 20-Oxo-leukotriene E4, and phenol sulfate, etc, in exposed zebrafish, and the shared significantly enriched calcium signaling pathways and necroptosis. The correlation coefficient was ≥ 0.6 and ≤−0.6, and the p-value was <0.05 between the control and exposure groups. |