Fig. 2
- ID
- ZDB-FIG-050825-3
- Publication
- Formstone et al., 2005 - Combinatorial activity of Flamingo proteins directs convergence and extension within the early zebrafish embryo via the planar cell polarity pathway
- Other Figures
- All Figure Page
- Back to All Figure Page
|
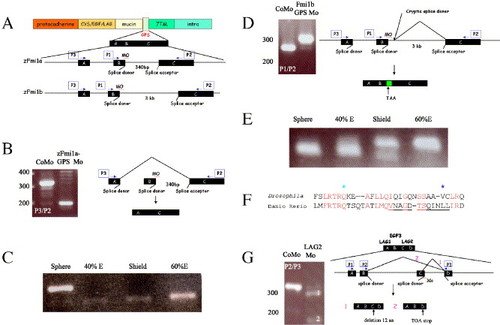
Generation of splicing defects in the GPS and LamG2 domains of Flamingo proteins. (A) Schematic representation of the Flamingo1 protein and morpholino-induced mis-splicing of the GPS domain. The GPS domain (pale green box) is encoded by three exons: A, B, and C. Position of morpholinos (MO) designed against targeted donor splice sites are indicated. Two sets of PCR using primer combinations P1/P2 and P3/P2 (blue arrows in cartoon) were used to assess mis-splicing. Abbreviations: Cys/EGF/LAG, cysteine-rich, EGF, and LamG2 domains; mucin is the serine/threonine-rich region; 7TM, seven-pass transmembrane domain; intra, intracellular domain. (B and C) Analysis of defective splicing in F1a-GPS1 morphants. (B) RT-PCR with primers P3 and P2 (blue arrows). Amplification of a smaller RT-PCR fragment (200 bp) than from controls (320 bp) was observed. CoMo is F1a-GPS1 morpholino but with four non-clustered, single nucleotide changes. Molecular weight markers (bp) are indicated to the left. Due to exon skipping, primers P1/P2 failed to amplify a band. Schematic representation of exon skipping induced by F1a-GPS is shown to the right. (C) Timing of onset of zFmi1a GPS domain splicing defect during embryogenesis. E is epiboly. (D and E) Analysis of defective splicing generated by injection of F1b-GPS1 morphants. (D) RT-PCR with primers P1, P2 is shown (blue arrows), primers P3/P2 amplified similar pattern of bands. RT-PCR of F1b-GPS1 morphant embryos amplified a larger band (310 bp) than controls (280 bp) due to use of a cryptic splice site that incorporated an extra 40 bp of intronic sequence into zFmi1b mRNA (indicated by green box). CoMo is F1b-GPS1 morpholino but with four nucleotide changes. Molecular weight markers (bp) are indicated to left of each gel. Schematic representation is shown to the right. (E) Timing of onset of the zFmi1b GPS domain splicing defect during embryogenesis, E is epiboly. (F and G) Imitation of Drosophila Flamingo PCP mutations using morpholino-induced mis-splicing. (F) Comparison of amino acid sequence from LamG2 in Drosophila melanogaster Flamingo and zFmi1a. Genomic sequence for zFmi1a was derived from ZFIN. Conserved amino acids are highlighted in red. Position of Drosophila mutation, Q1838 to stop, is marked by a pale blue star and mutation, V1849 to D, is marked by a dark blue star. 12 amino acid deletion induced by LamG2 morpholino is underlined. (G) Analysis of defective splicing generated by injection of F1a-LamG2 morpholino into zebrafish embryos. RT-PCR analysis utilized primers P3 and P2 (blue arrows), primers P1, P3 amplified a similar pattern of bands. Molecular weight markers (bp) are indicated to left of gel. Schematic representation of defective splicing within zFmi1a LamG2 domains induced by injection of LamG2 morpholino is shown to the right. The LAG1/EGF3/LAG2 domain, part of the pale orange box in (A), is encoded by four exons, A encodes C-terminal of LamG1, B encodes EGF3, C plus D encode N-terminal part of LamG2. Two mis-spliced products were resolved on the gel: (1) indicates the largest and most predominant of these resulting from use of a cryptic splice site within exon C, 36 bp upstream of the original acceptor site that deleted 12 amino acids from the LamG2 domain; (2) indicates smaller and less abundant product resulting from exon-skipping event that truncates the protein within the LamG2 domain. |
| Genes: | |
|---|---|
| Fish: | |
| Knockdown Reagents: | |
| Anatomical Term: | |
| Stage Range: | Sphere to Shield |
Reprinted from Developmental Biology, 282(2), Formstone, C.J., and Mason, I., Combinatorial activity of Flamingo proteins directs convergence and extension within the early zebrafish embryo via the planar cell polarity pathway, 320-335, Copyright (2005) with permission from Elsevier. Full text @ Dev. Biol.