- Title
-
Active receptor tyrosine kinases, but not Brachyury, are sufficient to trigger chordoma in zebrafish
- Authors
- D'Agati, G., Cabello, E.M., Frontzek, K., Rushing, E.J., Klemm, R., Robinson, M.D., White, R.M., Mosimann, C., Burger, A.
- Source
- Full text @ Dis. Model. Mech.
|
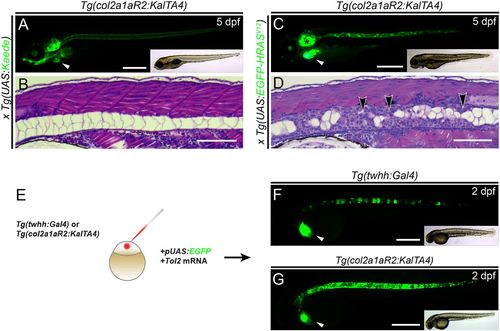
Tg(col2a1aR2:KalTA4) enables stable and transient oncogene expression in the developing notochord. (A,B) Lateral view (A) and transverse histology section (stained with H&E) (B) of 5 dpf zebrafish embryos transgenic for the transgene col2a1aR2:KalTA4 and crossed with stable UAS:Kaede (visible as green fluorescence in A). Note expression in the notochord, craniofacial cartilage, otic vesicle and pectoral fins; the prominent green heart (white arrowhead, A) indicates the myl7:EGFP transgenesis marker associated with col2a1aR2:KalTA4. The developing notochord shows that the large vacuolated cells take up the vast majority of the notochord volume and are rimmed by a thin layer of sheath cells (B). (C,D) Expression of stable UAS:EGFP-HRASV12 (detectable by green fluorescence of the fusion protein, C) by col2a1aR2:KalTA4causes invasive and widespread notochord hyperplasia (black arrowheads, D) and overgrowth of other cartilage tissue (i.e. otic vesicle, black asterisk in C); the white arrowhead indicates the myl7:EGFP transgenesis marker (A). (E-G) col2a1aR2 provides a potent driver for transient notochord expression. (E) Injection of UAS:EGFP into the one-cell-stage embryos with either twhh:Gal4 (F) or col2a1aR2:KalTA4 (G) to visualize the notochord mosaicism resulting from random integration of UAS:EGFP by Tol2 transposase. While injections into twhh:Gal4 result in highly patchy EGFP expression (F, green fluorescence; n=33/56), col2a1aR2:KalTA4 more consistently drives EGFP expression throughout the notochord (G, green fluorescence; n=25/47); n indicates representative EGFP-expressing embryos in an injected representative clutch. Scale bars: 500 μm in A,C,F,G; 200 μm in B,D. See also Fig. S1. |
|
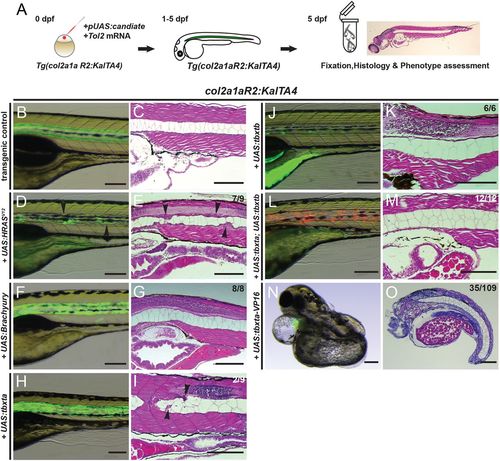
Overexpression of brachyury genes in the zebrafish notochord is insufficient to initiate chordoma. (A) Workflow of injection-based notochord hyperplasia assessment: at the one-cell stage, Tg(col2a1aR2:KalTA4) embryos are injected with Tol2 transposase mRNA and a plasmid containing a fluorescently labeled candidate gene under UAS control; injected embryos are raised up to 5 dpf and candidate gene expression is monitored through notochord fluorescence. Embryos with consistent reporter expression are fixed, sectioned and stained with H&E to assess the notochord phenotype using light microscopy. (B-O) Close-up lateral view of embryo notochords at 5 dpf, for brightfield and fluorescence (left column) and H&E histology (right column; different embryos per condition); numbers indicate observed versus total from an individual representative experiment. (B,C) The col2a1aR2:KalTA4 control reference at 5 dpf, expressing UAS:Kaede to fluorescently label the notochord. (D,E) Transient injection of UAS:EGFP-HRASV12 causes localized notochord hyperplasia (arrowheads). (F-I) Forced expression of human TBXT (brachyury) (F,G) or of the zebrafish gene tbxta (H,I) does not affect notochord development or cell proliferation. Minor lesions caused by collapsed vacuolated cells developed in a few UAS:tbxta-expressing notochords (arrowheads, I). (J,K) Overexpression of the second zebrafish brachyury gene tbxtb does not affect notochord development or proliferation. (L,M) Combined overexpression of both zebrafish brachyury genes tbxta and tbxtb has no effect on proliferation and the notochord develops normally. (N,O) Notochord-driven expression of tbxta-VP16, encoding a dominant-active transcriptional activator, leads to non-autonomous defects in the trunk with a shortened and/or severely curled trunk (32% of analyzed embryos), whereas the notochord forms normally. Scale bars: 200 μm. |
|
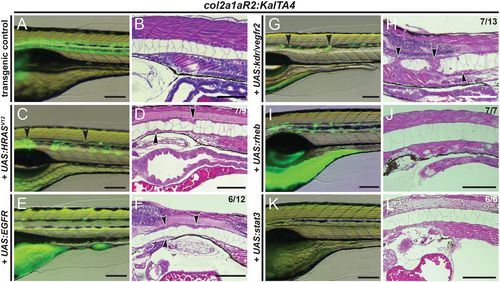
Transient overexpression of RTK genes drives notochord hyperplasia. (A-L) Close-up lateral views of embryo notochords at 5 dpf, with brightfield/fluorescence (left panels) and H&E histology (right panels; different embryos per condition); numbers in D,F,H and J indicate embryos with observed lesions versus phenotypically normal embryos observed in an individual representative experiment, and numbers in L indicate wild-type-looking embryos (n=3 experiments). (A,B) The col2a1aR2:KalTA4 control reference at 5 dpf, expressing UAS:Kaede to fluorescently label the notochord. (C,D) Transient injection of UAS:EGFP-HRASV12 causes localized hyperplasia (arrowheads, C,D) in the notochord. (E,F) Overexpression of human EGFRconsistently causes local hyperplasia in the developing notochord (arrowheads, F). (G,H) Overexpression of zebrafish kdr (encoding the VEGFR2 ortholog) causes strong hyperplasia (arrowheads in G,H, compare with D,F). (I,J) Zebrafish rheb overexpression leads to enlarged vacuoles and no detectable hyperplasia. (K,L) Zebrafish stat3 overexpression leads to no detectable hyperplasia and allows for normal notochord development within 5 dpf. Scale bars: 200 μm. |
|
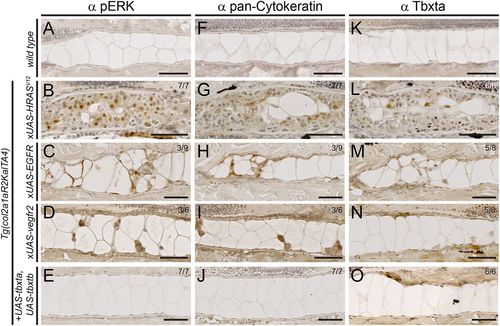
Expression of chordoma markers in RTK-transformed zebrafish notochords. (A-O) Immunohistochemistry on sagittal sections through the notochord of 5 dpf zebrafish embryos of the indicated genotypes, expressing either stable or mosaic transgenes. (A-E) MAPK pathway activation through HRASV12, EGFR and kdr overexpression results in nuclear pERK staining in the notochord, whereas controls and tbxta,tbxtb-injected embryos are negative for pERK staining. (F-J) HRASV12, EGFR and kdr overexpression results in staining for pan-Cytokeratin, whereas tbxta,tbxtb-injected embryos are negative for pan-Cytokeratin staining akin to wild-type controls. (K-O) Whereas control notochords show faint to no Tbxta signal, owing to low cell density of the sheath layer (K), HRASV12-overexpressing notochords (L) as well as EGFR-overexpressing (M) and kdr-overexpressing (N) notochords show prominent nuclear Tbxta staining. The tbxta,tbxtb-overexpressing notochords also stain positive (O), confirming transgenic tbxta expression. Scale bars: 50 µm |

ZFIN is incorporating published figure images and captions as part of an ongoing project. Figures from some publications have not yet been curated, or are not available for display because of copyright restrictions. PHENOTYPE:
|
|
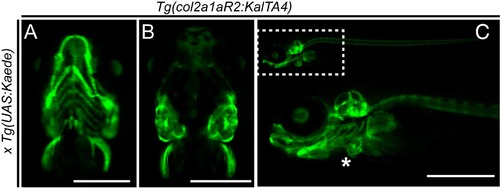
Activity of Tg(col2a1a:KalTA4) at later developmental stages and adults (A-C) col2a1aR2:Kal4TA4 activity shown by crossed-in UAS:Kaede expression (green fluorescence). (A) Ventral view showing the developing jaw cartilage and pectoral fins at 5 dpf. (B) Dorsal view showing activity in otic vesicle cartilage and pectoral fins at 5 dpf. (C) Lateral view of the craniofacial cartilage expressing col2a1aR2:KalTA4 at 5 dpf. Asterisk marks the heart labelled with myl7:EGFP as transgenesis marker. See also Figure 1 EXPRESSION / LABELING:
|
|
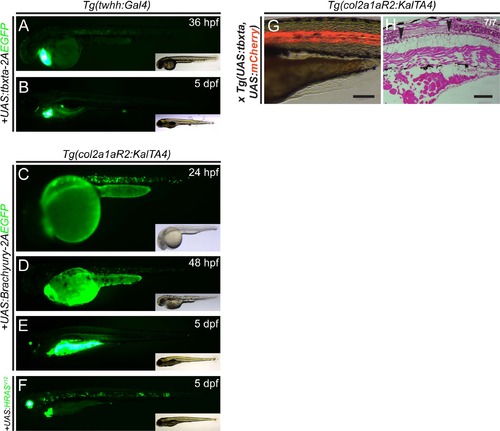
Tagged fusion proteins of Brachyury fail to express efficiently in vivo. (A-F) Lateral views of zebrafish embryos at indicated developmental stages, with EGFP fluorescence (green) in main panels, insets show brightfield for overall embryo morphology. (A,B) Transgene silencing in Tg(twhh:Gal4) with injected UAS:tbxta-2A-EGFP; while selectively expressed in the notochord at 36 hpf (n= 43/56), expression greatly reduced at 5 dpf. (C-F) Representative embryos transiently expressing UAS: Brachyury-2A-EGFP at 24 hpf at detectable levels (C), while expression becomes significantly reduced at 48 hpf (D) and is completely silenced at 5 dpf (E). (F) Transiently expressed UAS:EGFPHRASV12 at 5 dpf mainly localizes in the notochord and to a minor extent in the otic vesicle. Tumorigenic lesions could only be detected in the notochord. Transgenic markers used are myl7:EGFP for twhh:Gal4 and col2a1aR2:KalTA4, cryaa:Venus for injected UAS-plasmids. (G,H) Akin to injected UAS:tbxta, stable transgenic Tg(UAS:tbxta) overexpression fails to cause any significant notochord defect, with only few areas showing smaller vacuolated cells (arrowheads, H). |
|
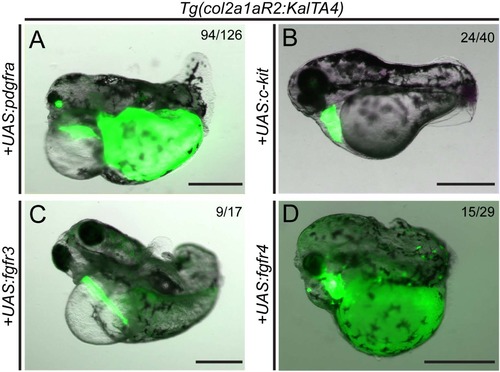
Embryo-wide, non-autonomous perturbation of embryo development upon notochord-specific overexpression of individual developmental RTK genes. (A-D) Representative Tg(col2a1aR2:KalTA4) embryos injected with individual UAS constructs for different chordoma-implicated RTK genes; numbers depict observed phenotype versus phenotypically normalappearing embryos in a representative experiment (n=3). Notochord specific overexpression of zebrafish pdgfra (A), c-kit (B), fgfr3 (C), and fgfr4 (D) leads to severe body axis truncation and aberrant development of cardiovascular and craniofacial structures (A, C, D: 5 dpf, B: 3 dpf). Transgenic markers: myl7:EGFP for col2a1aR2:KalTA4, cryaa:Venus for injected UAS plasmids. Scale: 500 μm. See also Figure 3. |