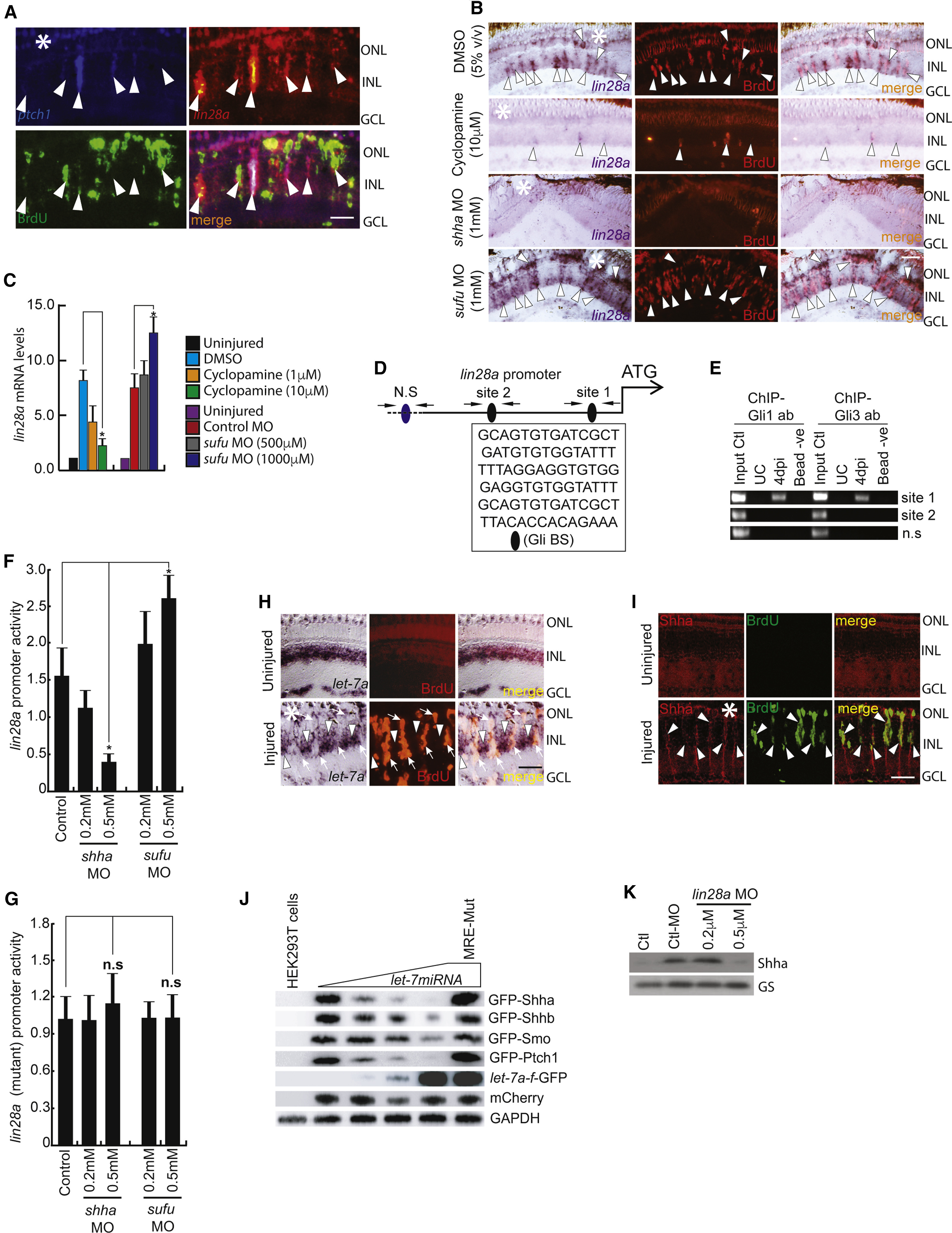
Fig. 3
Lin28a-let-7 Axis Regulates Shh Signaling Component Genes in the Injured Retina
(A) FISH and IF microscopy images of a 0.5-μm-thick optical section of retina showed co-localization of lin28a with ptch1 in BrdU+ MGPCs at 4 dpi. Arrowheads mark co-expression of genes in BrdU+ cells.
(B and C) BF microscopy images of lin28a mRNA ISH in the retina at 4 dpi with cyclopamine treatment and shha or sufu knockdown (B), which was quantified by qPCR (C). Arrowheads mark co-expression of genes in BrdU+ cells in (B).
(D and E) Schematic of the lin28a promoter with a potential Gli-BS cluster, where arrows mark ChIP primers and capital letters mark consensus sequence of Gli-BS (D). A 4 dpi retinal ChIP assay showed both Gli1 and Gli3 bound to one of the two Gli-BS clusters (E).
(F) Luciferase assay in 24 hpf embryos co-injected with lin28a:GFP-luciferase vector and sufu or shha MOs.
(G) Luciferase assay with mutated Gli-BSs of the lin28a promoter in an experiment similar to (F).
(H and I) ISH and IF microscopy of retina showing co-exclusion of let-7a microRNA (H) and co-localization of Shha protein (I) in BrdU+ MGPCs in the retina at 4 dpi. Arrowheads mark expression of let-7a in BrdU− cells and arrows mark co-exclusion of let-7a from BrdU+ cells in (H). Arrowheads mark co-expression of Shha in BrdU+ cells in (I).
(J) let-7 microRNA downregulated the translation of GFP fused with the indicated gene constructs harboring microRNA-binding regions in a dose-dependent manner in HEK293T cells.
(K) Western blot of Shha in lin28a-MO electroporated retina at 4 dpi.
Scale bars represent 10 μm (A, H, and I) and 20 μm (B). Asterisk indicates the injury site (A, B, H, and I). Error bars represent SD.∗p < 0.001 (C); ∗p < 0.001 (F). n = 6 biological replicates (C, F, and G). GS, glutamine synthetase. See also Figures S3, S6, and S7.