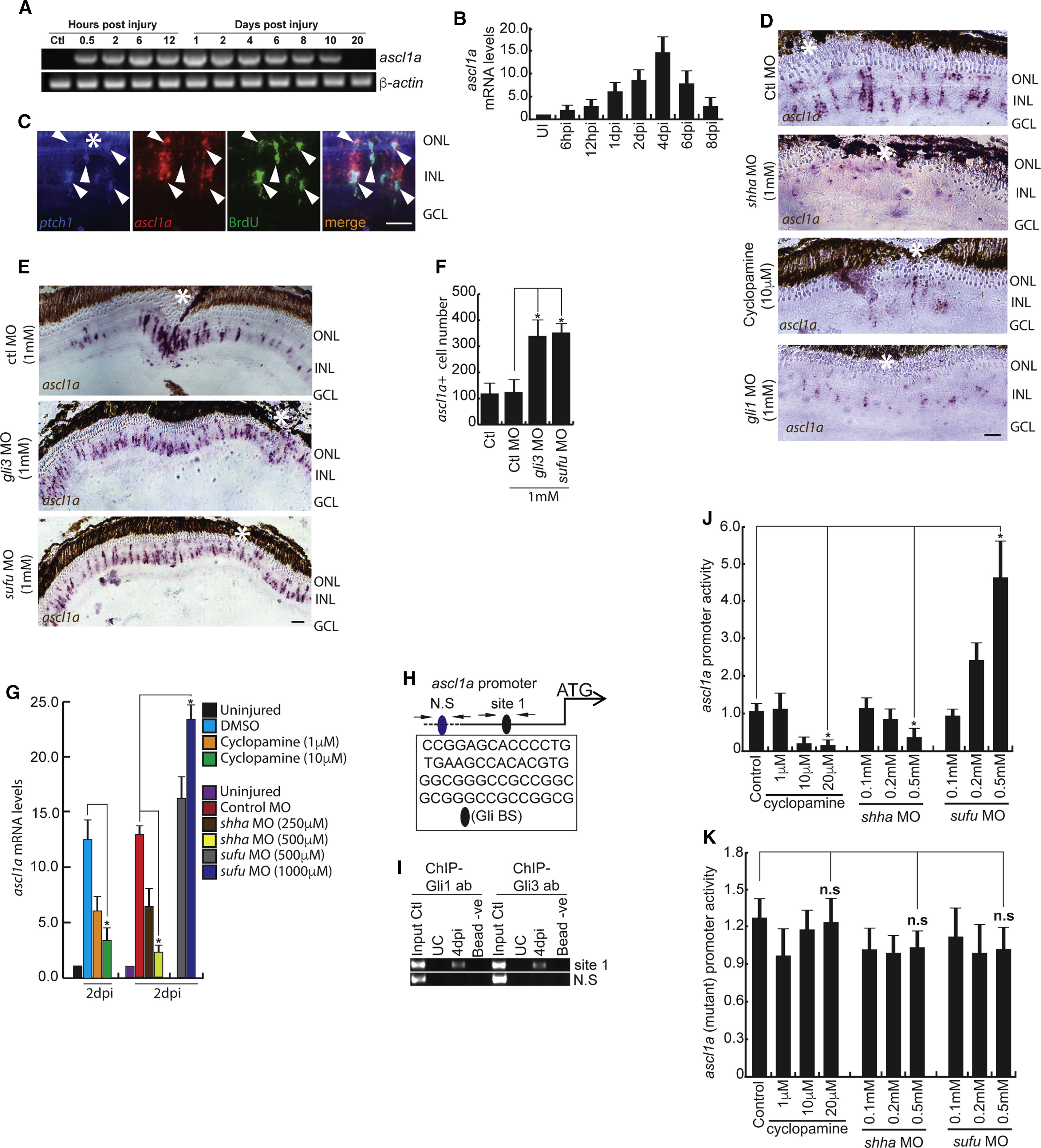
Fig. 2
Shh-Signaling-Dependent ascl1a Regulation in the Injured Retina
(A and B) RT-PCR (A) and qPCR (B) analysis of ascl1a in the post-injured retina; n = 6 biological replicates.
(C) Fluorescence ISH (FISH) and IF microscopy images of a 0.5-μm-thick optical section of retina showing co-localization of ascl1a with ptch1 in BrdU+ MGPCs at 4 dpi. Arrowheads mark co-expression of genes in BrdU+ cells.
(D–F) BF microscopy images of ascl1a mRNA ISH in retina at 4 dpi with cyclopamine treatment, shha or gli1 knockdowns (D), and gli3 or sufu knockdowns (E). The number of ascl1a+ cells from (E) is quantified in (F).
(G) qPCR analysis of ascl1a mRNA with cyclopamine treatment and shha or sufu knockdown in 2 dpi retina.
(H) Schematic of the ascl1a promoter with a putative Gli-binding site (Gli-BS) cluster. Arrows mark ChIP primers, N.S marks the negative control, and capital letters mark putative Gli-BSs.
(I) Retinal ChIP assay at 4 dpi showing both Gli1 and Gli3 bound to the ascl1a promoter.
(J) Luciferase assay in 24 hpf embryos co-injected with ascl1a:GFP-luciferase vector and sufu or shha MOs.
(K) Luciferase assay was done with mutated Gli-BS of ascl1a promoter in an experiment similar to (J).
Scale bars represent 10 μm in (C) and 20 μm in (D) and (E). Asterisk indicates the injury site (C–E). Error bars represent SD. ∗p < 0.0001 (F); ∗p < 0.01 (G); ∗p < 0.01 (J). n = 6 biological replicates (F and G); n = 3 (J). See also Figures S2, S6, and S7.