- Title
-
NPC1 links cholesterol trafficking to microglial morphology via the gastrosome
- Authors
- Zareba, J., Cattaneo, E.F., Villani, A., Othman, A., Streb, S., Peri, F.
- Source
- Full text @ Nat. Commun.
|
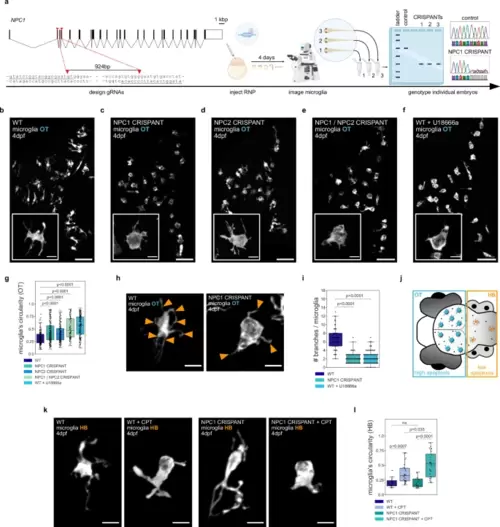
NPC1 and NPC2 deficiencies lead to ameboid microglia under phagocytic stress.a Schematic of the CRISPANT workflow created in BioRender. Zareba, J. (2022) BioRender.com/v30d049. b–f Representative images of OT zebrafish microglia (Tg(mpeg1.1:EGFP-CAAX)). Scale bar 50 µm. Boxes with magnifications of single microglia (TgBAC(csf1ra:GAL4-VP16), Tg(UAS-E1B:NTR-mCherry)). Scale bar 10 µm. Wild-type (b), NPC1 CRISPANT (c), NPC2 CRISPANT (d), NPC1 / NPC2 CRISPANT (e), wild-type treated with U18666a (f). g Circularity of OT microglia in wild-type (N = 10, n = 207), NPC1 CRISPANT (N = 14, n = 230), NPC2 CRISPANT (N = 10, n = 193), NPC1 / NPC2 CRISPANT (N = 6, n = 115) and wild-type zebrafish treated with U18666a (N = 10, n = 223). h Representative images of OT zebrafish microglia (TgBAC(csf1ra:GAL4-VP16), Tg(UAS-E1B:NTR-mCherry)). Branches indicated by orange arrowheads. Scale bar 10 µm. i Number of branches per OT microglia from wild-type (N = 3, n = 54), NPC1 CRISPANT (N = 6, n = 85) and wild-type treated with U18666a (N = 4, n = 95) zebrafish. j Schematic of the zebrafish brain regions. k Representative images of HB zebrafish microglia (TgBAC(csf1ra:GAL4-VP16), Tg(UAS-E1B:NTR-mCherry)). Scale bar 10 µm. l Circularity of HB zebrafish microglia from wild-type (N = 8, n = 16), wild-type treated with CPT (N = 15, n = 28), NPC1 CRISPANT (N = 7, n = 16) and NPC1 CRISPANT treated with CPT (N = 12, n = 28). N refers to the number of zebrafish embryos and n to the number of microglia examined. Boxplots represent the median value and interquartile range; the ends of the whiskers correspond to the minimum and maximum values. Statistical tests: Mann-Whitney-Wilcoxon test two-sided with Bonferroni correction. ns without any additional p value on the graph stands for p = 1. OT stands for optic tectum and HB stands for hindbrain. Source data are provided as a Source Data file. EXPRESSION / LABELING:
PHENOTYPE:
|
|
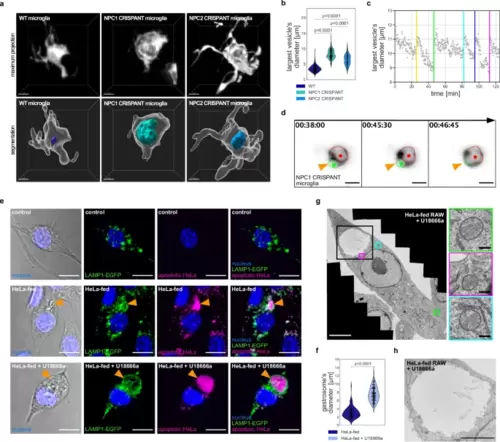
NPC1 deficiency causes expansion of the gastrosome in phagocytic cells.a Representative images of OT microglia (TgBAC(csf1ra:GAL4-VP16), Tg(UAS-E1B:NTR-mCherry)) in the zebrafish displayed as maximum projections and segmented images. The largest vesicle present in each cell is displayed as a segmentation. Scale bar 5 µm. b Diameter of the largest vesicle in wild-type (N = 26, n = 102), NPC1 CRISPANT (N = 29, n = 232) and NPC2 CRISPANT (N = 7, n = 126) OT microglia. N refers to the number of zebrafish embryos and n to the number of microglia examined. c Graph representing the diameter of the largest vesicle found in NPC1 CRISPANT OT microglia over time. Colored vertical lines indicate fusion with incoming phagosomes. d Representative images of fusion events between the large vesicle (marked with red) and incoming phagosome (marked with green) in NPC1 CRISPANT OT microglia (TgBAC(csf1ra:GAL4-VP16), Tg(UAS-E1B:NTR-mCherry)). Scale bar 10 µm. Related to Supplementary Movie 3. e Representative images of RAW LAMP1-EGFP macrophages. Orange arrowhead indicates the gastrosome. Scale bar 10 µm. f Diameter of the gastrosome in HeLa-fed RAW LAMP1-EGFP macrophages, untreated (n = 69) and treated with U18666a (n = 115). n refers to the number of cells examined. g EM image of HeLa-fed RAW treated with U18666a. Lysosomes are framed in colored squares and the gastrosome is framed in the black square (enlargement in Fig. 3h). Scale bar 5 µm. Enlarged regions with lysosomes on the right. Scale bar 0.25 µm. h Electron microscope image of the gastrosome in HeLa-fed RAW treated with U18666a. Scale bar 3 µm. Statistical tests: Mann-Whitney-Wilcoxon test two-sided with Bonferroni correction. OT stands for optic tectum. Source data are provided as a Source Data file. |
|
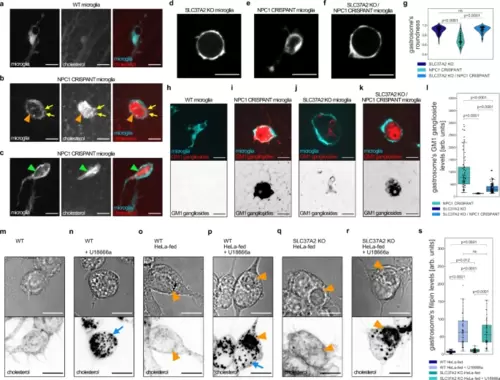
Gastrosome in NPC1 deficient cells accumulates cholesterol and GM1 gangliosides.a–c Representative images of OT microglia (TgBAC(csf1ra:GAL4-VP16), Tg(UAS-E1B:NTR-mCherry)) with TopFluor cholesterol. Scale bar 10 µm. b Orange arrowhead indicates gastrosome and yellow arrows point to smaller cholesterol-rich vesicles. c Green arrowhead points to a visible cholesterol crystal. d–f Representative images of OT microglia (TgBAC(csf1ra:GAL4-VP16), Tg(UAS-E1B:NTR-mCherry)). Scale bar 20 µm. g Roundness of the gastrosome in SLC37A2 KO (N = 3, n = 69), NPC1 CRISPANT (N = 4, n = 59) and NPC1 CRISPANT in SLC37A2 KO background (N = 4, n = 61) in OT microglia (TgBAC(csf1ra:GAL4-VP16), Tg(UAS-E1B:NTR-mCherry)). N refers to the number of zebrafish embryos and n to the number of microglia examined. h–k, Representative images of OT microglia (TgBAC(csf1ra:GAL4-VP16), Tg(UAS-E1B:NTR-mCherry)) fixed and stained for GM1 gangliosides. Scale bar 10 µm. l GM1 ganglioside levels in the gastrosome from NPC1 CRISPANT (N = 9, n = 68), SLC37A2 KO (N = 7, n = 69) and NPC1 CRISPANT in SLC37A2 KO background (N = 9, n = 120) fixed OT microglia. N refers to the number of zebrafish embryos and n to the number of microglia examined. m–r Representative images of RAW macrophages stained with filipin. Scale bar 10 µm. Blue arrows point to cholesterol-rich small vesicles while the orange arrowheads indicate the gastrosome. s Filipin levels in the gastrosome of RAW macrophages: WT HeLa-fed (n = 40), WT HeLa-fed and U18666a treated (n = 54), SLC37A2 KO HeLa-fed (n = 30), SLC37A2 KO HeLa-fed and U18666a treated (n = 50). n refers to the number of cells examined. Boxplots represent the median value and interquartile range; the ends of the whiskers correspond to the minimum and maximum values. Statistical tests: Mann-Whitney-Wilcoxon test two-sided with Bonferroni correction. ns without any additional p-value on the graph stands for p = 1. OT stands for optic tectum. Source data are provided as a Source Data file. PHENOTYPE:
|
|
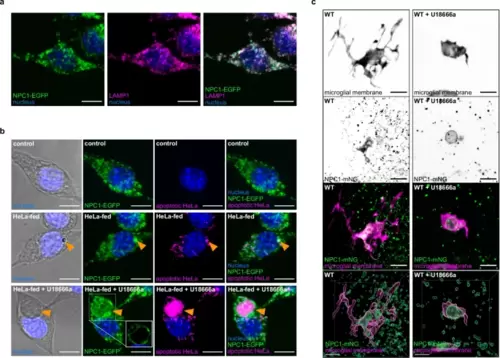
NPC1 localizes on the gastrosome that accumulates apoptotic material.a Representative image of RAW NPC1-EGFP macrophages stained for LAMP1 (N = 2, more than 4 cells examined per condition in each repetition). Scale bar 10 µm. b Representative images of RAW NPC1-EGFP macrophages from in vitro feeding experiment. Gastrosome indicated by orange arrowhead. For HeLa-fed U18666a treated example, single plane view shown in a box for NPC1-EGFP (N = 2, more than 4 cells examined per condition in each repetition). Scale bar 10 µm. c, Representative images of OT (TgBAC(csf1ra:GAL4-VP16), Tg(UAS-E1B:NTR-mCherry; npc1:mNeonGreen)) zebrafish microglia showing localization of NPC1 displayed as maximum projection and segmentation (N = 4). Scale bar 10 µm. N refers to the number of experimental replicates. OT stands for optic tectum. |
|
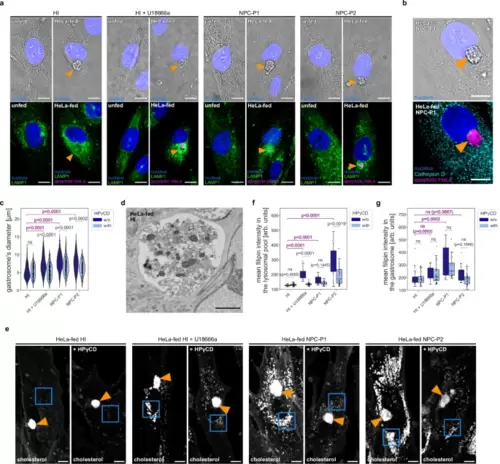
Patient-derived fibroblasts accumulate apoptotic material in the gastrosome that is larger in NPC deficient cases.a Representative images of human patients’ fibroblasts stained for LAMP1. The gastrosome is indicated with an orange arrowhead. Scale bar 10 µm. b Representative image of HeLa-fed human patients’ fibroblasts stained for Cathepsin D. The gastrosome is indicated with an orange arrowhead. Scale bar 10 µm. c Diameter of the gastrosome in HeLa-fed human patients’ fibroblasts: HI (n = 114), HI treated with HPγCD (n = 113), HI treated with U18666a (n = 126), HI treated with U18666a and HPγCD (n = 176), NPC-P1 (n = 156), NPC-P1 treated with HPγCD (n = 146), NPC-P2 (n = 113), NPC-P2 treated with HPγCD (n = 94). d Representative EM image of the gastrosome in HeLa-fed human patients’ fibroblast. Scale bar 3 µm. e Representative images of HeLa-fed human patients’ fibroblast stained with filipin. The gastrosome is indicated with an orange arrowhead and lysosomal pools are framed in blue squares. Scale bar 10 µm. f, g Mean intensities of the filipin staining from HeLa-fed human patients’ fibroblasts: HI (n = 25), HI treated with HPγCD (n = 30), HI treated with U18666a (n = 16), HI treated with U18666a and HPγCD (n = 32), NPC-P1 (n = 21), NPC-P1 treated with HPγCD (n = 21), NPC-P2 (n = 23), NPC-P2 treated with HPγCD (n = 33) in the lysosomal pool (framed in blue squares in Fig. 5e) (f), in the gastrosome (indicated with an orange arrowhead in Fig. 5e) (g). n refers to the number of cells examined. Boxplots represent the median value and interquartile range; the ends of the whiskers correspond to the minimum and maximum values. Statistical tests: Mann-Whitney-Wilcoxon test two-sided with Bonferroni correction. ns without any additional p-value on the graph stands for p = 1. HI stands for healthy individual. NPC1-P1 corresponds to GM18393 NPC patient and NPC1-P2 corresponds to GM17919 NPC patient. Source data are provided as a Source Data file. |
|
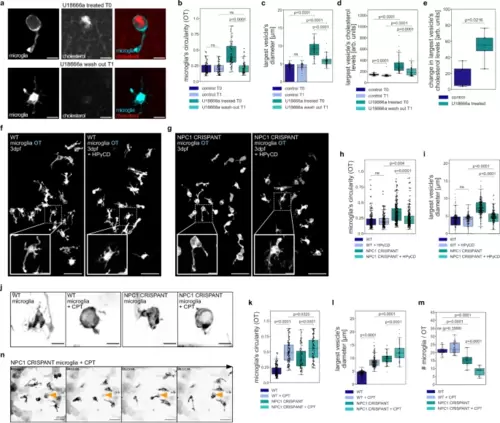
NPC deficient microglia are sensitive to elevated neuronal cell death.a Representative images from U18666a treated (T0) OT microglia (TgBAC(csf1ra:GAL4-VP16), Tg(UAS-E1B:NTR-mCherry)) and from wash out (T1) with TopFluor cholesterol. Scale bar 10 µm. b–e Measurements from control T0 (N = 5, n = 76), control T1 (N = 5, n = 76), U18666a treated T0 (N = 5, n = 71) and U18666a wash out T0 (N = 5, n = 71) zebrafish with TopFluor cholesterol of OT microglia circularity (b), quantification of the largest vesicle’s diameter (c), quantification of the largest’s vesicle cholesterol levels (d) and its changes per individual zebrafish brains (e). f, g Representative images of zebrafish OT microglia (TgBAC(csf1ra:GAL4-VP16), Tg(UAS-E1B:NTR-mCherry)) from HPγCD experiment. Scale bar 50 µm. Examples of single microglia in boxes. Scale bar 10 µm. h, i Measurements from wild-type (N = 10, n = 201), wild-type treated with HPγCD (N = 10, n = 191), NPC1 CRISPANT (N = 18, n = 291) and NPC1 CRISPANT treated with HPγCD (N = 18, n = 291) zebrafish of OT microglia circularity (h) and quantification of the largest vesicle’s diameter (i). j Representative images of OT microglia (TgBAC(csf1ra:GAL4-VP16), Tg(UAS-E1B:NTR-mCherry)) from Camptothecin (+ CPT) experiment. Scale bar 10 µm. k Circularity of OT microglia from wild-type (N = 10, n = 198), wild-type treated with CPT (N = 10, n = 123), NPC1 CRISPANT (N = 8, n = 84) and NPC1 CRISPANT treated with CPT (N = 15, n = 95) zebrafish. l Quantification of the largest vesicle’s diameter in wild-type (N = 10, n = 218), wild-type treated with CPT (N = 10, n = 221), NPC1 CRISPANT (N = 8, n = 80) and NPC1 CRISPANT treated with CPT (N = 9, n = 66) zebrafish OT microglia. m Number of OT microglia (TgBAC(csf1ra:GAL4-VP16), Tg(UAS-E1B:NTR-mCherry)) in wild-type (N = 23), wild-type treated with CPT (N = 31), NPC1 CRISPANT (N = 16), NPC1 CRISPANT treated with CPT (N = 15) zebrafish. n Representative images of OT microglia (TgBAC(csf1ra:GAL4-VP16), Tg(UAS-E1B:NTR-mCherry)) treated with CPT. Left side of Supplementary Movie 7. Dying cell and its subsequent engulfment indicated by orange arrowhead. Scale bar 40 µm. N refers to the number of zebrafish embryos and n to the number of microglia examined. Boxplots represent the median value and interquartile range; the ends of the whiskers correspond to the minimum and maximum values. Statistical tests: Mann-Whitney-Wilcoxon test two-sided with Bonferroni correction. ns without any additional p-value on the graph stands for p = 1. OT stands for optic tectum and CPT stands for Camptothecin. Source data are provided as a Source Data file. PHENOTYPE:
|