- Title
-
miRNA-7145-cuedc2 axis controls hematopoiesis through JAK1/STAT3 signaling pathway
- Authors
- Xu, Y., Guo, R., Huang, T., Guo, C.
- Source
- Full text @ Cell Death Discov
|
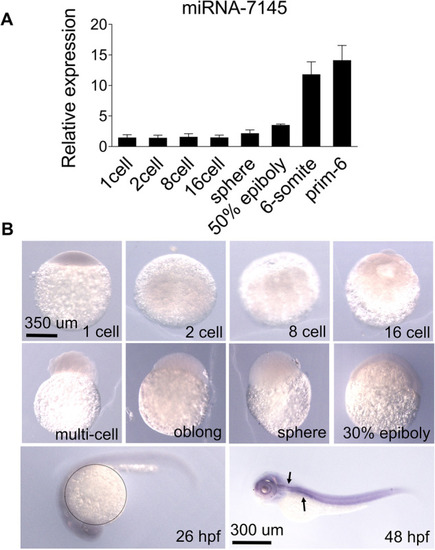
Spatial-temporal expression pattern of miRNA-7145 in wild-type embryos. |
|
Effect of miRNA-7145 overexpression on hematopoiesis. The embryos were injected with 200 pg miRNA-7145 mimics at the 1 cell stage, and the embryos were collected at 26 hpf. |
|
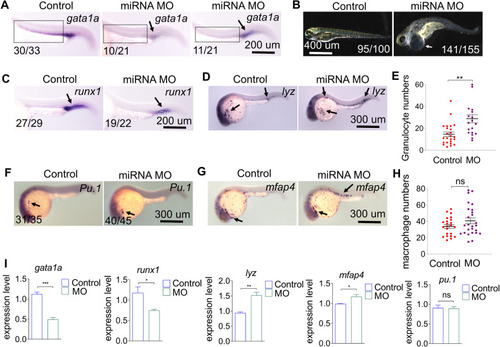
Effect of miRNA-7145 knock-down on hematopoiesis. Embryos were injected with 10 ng miRNA-7145 MO at the 1 cell stage, and the embryos were collected at 26 hpf and 4 dpf. |
|
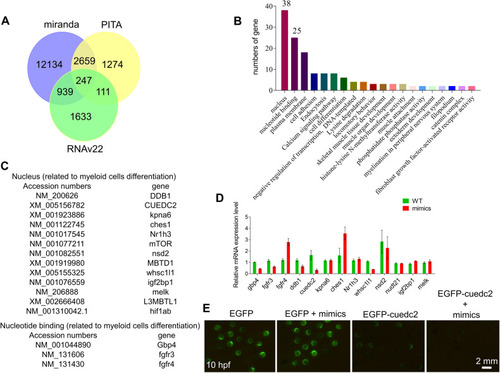
A target gene of miRNA-7145 was identified by bioinformatics analysis and fluorescence reporting experiments. |
|
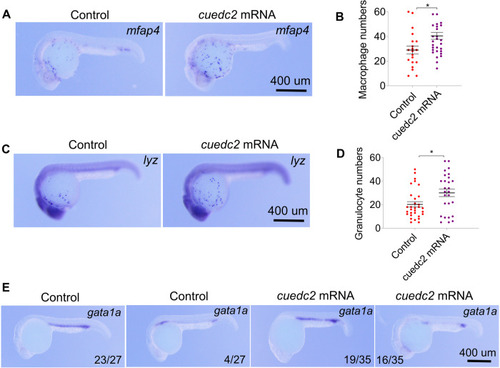
Effect of cuedc2 overexpression on hematopoiesis. Embryos were injected with 200 pg cuedc2 mRNA at the 1 cell stage, and the embryos were collected at 26 hpf. |
|
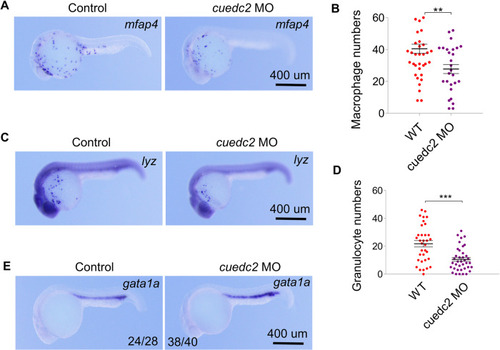
Effect of cuedc2 knock-down on hematopoiesis. The embryos were injected with 10 ng cuedc2 MO at the 1 cell stage, and the embryos were collected at 26 hpf. |
|
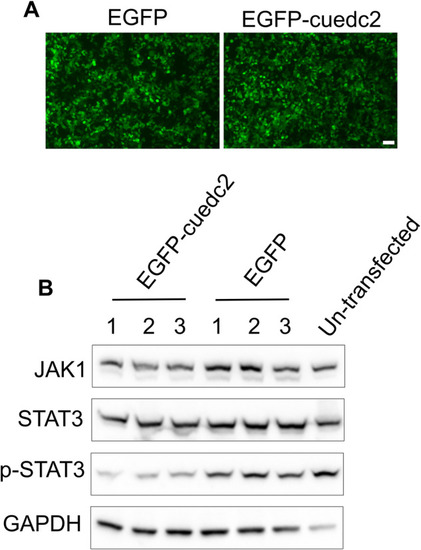
Effect of zebrafish cuedc2 overexpression on the JAK1/STAT3 signaling pathway. |