- Title
-
PML and PML-like exonucleases restrict retrotransposons in jawed vertebrates
- Authors
- Mathavarajah, S., Vergunst, K.L., Habib, E.B., Williams, S.K., He, R., Maliougina, M., Park, M., Salsman, J., Roy, S., Braasch, I., Roger, A.J., Langelaan, D.N., Dellaire, G.
- Source
- Full text @ Nucleic Acids Res.
|
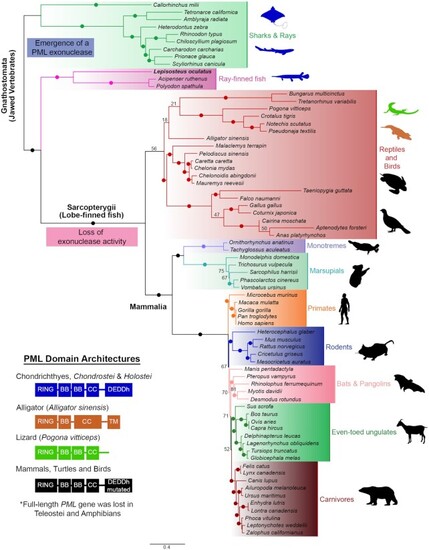
Domain architecture and Maximum Likelihood phylogenetic tree showing the relationships of PML homologs. 70+ protein sequences encoded in genomes from representative vertebrates were identified and utilized to construct the phylogenetic tree. PML emerged as an exonuclease in jawed vertebrates and eventually lost the catalytic residues for activity in amniote species. Teleost fishes and amphibians lack full-length PML-I. Ultrafast bootstrap values are shown as numbers if <85% and indicated with solid dots on branches if > 85%. Branch lengths indicate the expected number of amino acids substitution per site. Domains for homologs were determined using InterPro (https://www.ebi.ac.uk/interpro/) and four different domain architectures (left) were seen across analyzed sequences. Domain abbreviations refer to RBCC (Ring-B-box-Coiled Coil), BB (B-box), CC (Coiled coil), and DEDDh (3′-5′ DEDDh exonuclease). Sequences from species that were analyzed are shown in silhouettes and coloured according to the corresponding domain architecture on the left. Blue silhouettes indicate representative species with a PML ortholog encoding a predicted exonuclease domain. Silhouettes for species were obtained from PhyloPic (https://phylopic.org). |
|
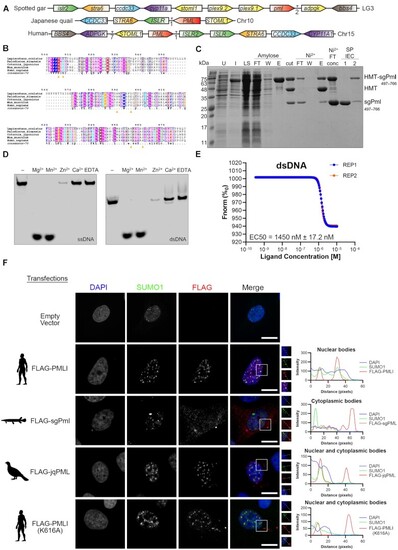
Spotted gar Pml is an active cytoplasmic DNA exonuclease. (A) Synteny of pml locus between the genomes of the Spotted gar, Japanese quail and Humans. Similarly coloured genes represent homologs between species. The intervening arrows with numbers indicate the number of adjacent genes between homologous genes. (B) MUSCLE alignment of the spotted gar PML (sgPml) C-terminal DEDDh exonuclease (CDE) domain protein with other PML orthologs. The consensus sequence is displayed below the sequences for residues with over 70% conservation across species. Yellow arrows indicate predicated catalytic residues for sgPml and the red circle indicates the conserved SUMOylation site (PML-I K616). (C) Purification of sgPml-CDE. sgPml-CDE was expressed as a fusion to sequences encoding for hexahistidine, maltose binding protein, and a TEV protease cleavage site (HMT). SDS-PAGE analysis is shown for samples of uninduced (U) and induced (I) cells, soluble lysate (LS), after TEV cleavage (cut), flowthrough (FT), wash (W), and elution (E) fractions of amylose and Ni2+ affinity chromatography, and fractions from ion exchange chromatography (IEC). (D) Exonuclease activity of sgPml-CDE requires Mg2+ or Mn2+. Fluorescent oligonucleotides were incubated with sgPml-CDE in the presence of the indicated divalent cation or EDTA. ‘-’ refers to input without the addition of cations. (E) sgPml binds to dsDNA with high affinity. Microscale thermophoresis was used to quantify the affinity between sgPml-CDE and dsDNA (n = 2). The different coloured lines are data shown for two replicates. (F) Subcellular localizations of FLAG-tagged PML-I, sgPml, quail (jqPML) and PML-I (K616A). Cells were transfected with either an empty vector or the indicated FLAG-tagged proteins. Colocalization between FLAG-tagged proteins and SUMO1 was assessed by immunostaining and shown at the right as line plot for the region of interest bound by the white box in the merged fluorescence image. Additional fields of view are shown in Supplementary Figure S16. Scale bars represent 10 μm. |
|
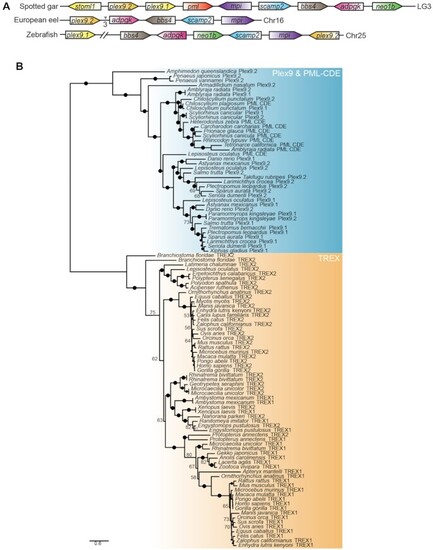
Origins of PML-like exon 9 (Plex9) and TREX1 genes. (A) Teleost plex9 genes show synteny to the spotted gar pml locus. Similarly coloured genes represent homologs between species. The chromosome where the pml locus is found in each species is indicated on the right. The intervening arrows with numbers indicate the number of intervening adjacent genes that exist between the homologous genes not found in the syntenic region. (B) Plex9 homologs resemble both TREX1/2 and PML exonucleases but are related to PML exonucleases. A maximum likelihood tree was constructed using Plex9, PML CDE and TREX1/2 protein sequences. While the TREX1 proteins cluster with TREX2, sequences for Plex9 cluster with PML CDE sequences. Bootstrap values are indicated beside each node. Ultrafast bootstrap values are shown as numbers if <85% and indicated with solid dots on branches if >85%. Branch lengths indicate the expected number of amino acid substitutions per site. |
|
Plex9 proteins represent a novel class of DEDDh DNA exonucleases. (A) Ectopic expression of zebrafish Plex9.1/.2 (zfPlex9.1/.2) and axTREX1/2 reveals that these proteins localize to the ER and cytoplasm. Proteins were FLAG-tagged and expressed in U2OS cells to assess their localization. A domain of Cytochrome p450 (CytoERM) tagged to mScarlet (addgene #85066) was utilized as an ER marker. An empty vector (BlueScript) was used as a control. Scale bars represent 10 μm. (B) The different domain architectures of TREX1/2 and Plex9 proteins and their known/determined localizations are shown on the right. TREX2 localization was assessed previously as both nuclear and cytoplasmic. (C) Purification of zfPlex9.1. zfPlex9.1 was expressed as a fusion to sequences encoding for hexahistidine, maltose binding protein, and a TEV protease cleavage site (HMT). SDS-PAGE analysis is shown for samples of uninduced (U) and induced (I) cells, soluble lysate (LS), after TEV cleavage (cut), flowthrough (FT), wash (W), and elution (E) fractions of amylose and Ni2+ affinity chromatography, and fractions from size exclusion chromatography (SEC). (D) zfPlex9.1 requires Mg2+ or Mn2+ for activity. Fluorescent oligonucleotides were incubated with zfPlex9 and the indicated metal. Samples were resolved by UREA-PAGE and visualized via fluorescence imaging. ‘-’ refers to input without the addition of cations. (E, F) DNA binding of zfPlex9.1. Microscale thermophoresis was used to quantify the affinity between zfPlex9.1 with (E) ssDNA and (F) dsDNA (n = 2). The different coloured lines are data shown for two replicates. |
|
sgPml, Plex9 and axTREX1 proteins suppress LINE-1 (L1) activity in an exonuclease-independent manner. (A) Retrotransposition of human L1 was assessed with the co-expression of the FLAG-tagged versions of the indicated proteins. Resultant plates from the assay (right) were stained with 0.5% crystal violet and quantified. Retrotransposition activity of zebrafish L1 elements were also assessed in (B, left) zfL2-1 and (C, right) zfL2-2 (n = 3 for human L1, zfL2-1 and zfL2-2). (D) A TREX1 knockout was generated using CRISPR/Cas9 and validated in U2OS (details in Supplementary Figure S12). (D) Retrotransposition activity was elevated in TREX1 knockouts and addback of the zfPlex9.1 and zfPlex9.2, as well as axTREX1 and axTREX2 (n = 3). (E) 2′,3′-cGAMP levels in U2OS WT and TREX1 knockout cells after the transfection of an empty vector and the human L1 vector (n = 3). 2′,3′-cGAMP concentrations were normalized to cell number for each experiment. All proteins were expressed with a FLAG epitope with the exception of GFP-TREX1(D18N). Controls represent empty vector (FLAG-tag backbone) used in the co-transfection with the L1 plasmid. All experiments were analysed by a repeated measures one-way ANOVA and then a Tukey's multiple comparisons test between groups. Asterisks indicate the following: ****P< 0.0001, ***P < 0.001, **P < 0.01, *P < 0.05. |
|
PML-I shuttles to the cytoplasm to suppress L1 by promoting the degradation of ORF1p. (A) PML mutants were generated encompassing the CDE (aa positions 596–882) and tagged with both an SV40 NLS and the PML-I NES at the N-terminus. Localizations of the mutants are shown in Supplementary Figure S14A. The mutants lack the C-terminal RBCC domain of PML. (B) Loss of PML elevates L1 activity which can be reversed with the addback of PML-I and the NES-PML mutant (n = 3). Resultant plates from the assay were stained with 0.5% crystal violet and quantified. Data was analysed by a repeated measures one-way ANOVA and then a Tukey's multiple comparisons test between groups. Asterisks indicate the following: ****P< 0.0001, ***P < 0.001, **P < 0.01, *P < 0.05. (C) Transfection of the human L1 retrotransposition vector led to cytoplasmic PML puncta forming. PML co-localized partly with TREX1 in the cytoplasm after L1 retroelements were active. (D) The nucleocytoplasmic shuttling of PML-I is CRM1-dependent. U2OS cells expressing GFP-PML-I were transfected with empty vector or L1 expression vector (L1.3, JM101.1) treated for 3 h with or without CRM1 inhibitor Leptomycin B (10 ng/ml). Cytoplasmic puncta of GFP-PML-I are indicated by the white the arrows. Scale bars represent 10 μm for images in (C) and (D). (E) U2OS cells (WT and PML KO) were transfected with an empty vector (-) or the L1 element with T7-tagged ORF1p. Cells were then treated with different concentrations of ubiquitin-proteasome inhibitor MG132 for 10 h and T7-ORF1p protein levels were examined. |
|
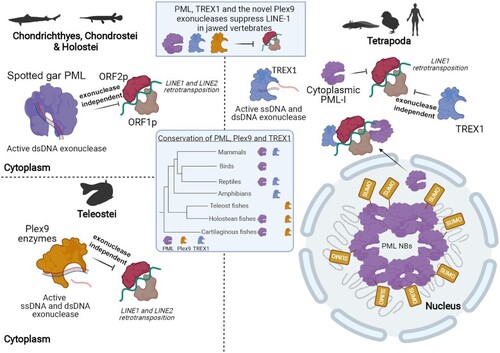
Overview of the suppression of LINE-1 retrotransposition in jawed vertebrates through the collective functions of PML, Plex9 and TREX1 enzymes. In fish, PML and novel Plex9 enzymes play a major role in suppressing both LINE-1 and LINE-2 elements. Despite the loss of PML in Teleost fish genomes, they have retained Plex9 genes for the purpose of restricting LINE retrotransposition. TREX1 appears later in tetrapods, where Plex9 genes are lost. In addition, enzymatic PML function is lost and the protein forms nuclear bodies rather than localizing to the cytoplasm. However, PML has a retained an exonuclease-independent role for the surveillance of LINEs by shuttling to the cytoplasm to suppress the LINE1 retrotransposition. Thus, the three protein families (PML, Plex9 and TREX) share an important function in maintaining genome integrity in jawed vertebrates by repressing LINE-1 propagation. |