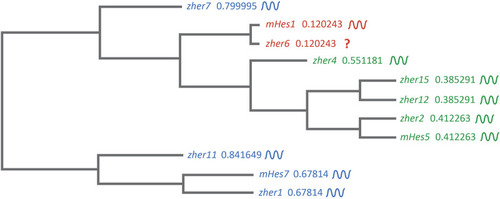
FIGURE 4
- ID
- ZDB-FIG-240213-12
- Publication
- Ramesh et al., 2024 - Species-specific roles of the Notch ligands, receptors, and targets orchestrating the signaling landscape of the segmentation clock
- Other Figures
- All Figure Page
- Back to All Figure Page
|
Clustal-based phylogenetic tree of mice (m) and zebrafish (z) |