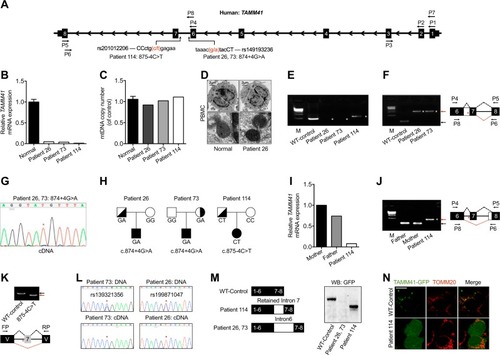
Variations of TAMM41 identified in AVSD patients. a Schematic diagram of TAMM41 transcript 005 (Ensemble ID: OTTHUMT00000339255) and the variations in Patient 26, 73, and 114 with their virtual positions in intron 6. b qPCR analysis of TAMM41 from whole blood cell extracted RNA with primers P2 and P3 (shown in a). The results show significantly reduced TAMM41 expression in the three affected children compared with six normal controls. c mtDNA copy number were evaluated by assessing the relative amounts of mitochondrial DNA and nuclear DNA (nDNA) of PBMCs from three patients and six healthy controls. MT-CO2 was used for as a marker for mtDNA and GAPDH for nDNA. d Representative TEM images showing mitochondrial morphology of PBMCs isolated from Patient 26 and one healthy control. Lower are enlarged ones showing mitochondria of the upper images. e Nest PCR with primers P1-P5 and P7-P6 (shown in a) failed to amplify full-length TAMM41 in Patient 26 and 73, and revealed an abnormally longer product in Patient 114. f Left is the DNA agarose gel electrophoresis showing that whilst normally spliced transcripts including exon 6, 7, and 8 (indicated by black arrows) amplified with primers P4 and P5 in control samples, the intron 6 retained transcript was detected via nested PCR (P4-P5 and P8-P6, indicated in a) in Patient 26 and 73. Right is the schematic drawing showing the different splicing patterns between the normal control (black dashed lines) and the patient samples (red dashed lines). g Sequencing of the PCR products in Patient 26, 73 (Fig. 1f) revealed the transcription of which from the mutant allele (indicated by red asterisks). h Transmission pattern of the TAMM41 variations identified in Patient 26, 73, and 114. i, j qPCR analysis indicating normal TAMM41 expression in sample from the father of Patient 114. The amplification products using P4-P5 and P8-P6 of Patient114 (red arrow), and her parents (black arrow) are shown on the left. Right is the schematic drawing indicating the different splicing events. k The variation (875-4C>T) in the splicing minigene reporter caused inactivation of the nearby splicing acceptor site and led to exon 7-skipping transcript generation. Lower is the schematic drawing manifesting the splicing pattern. l Exclusive expression from one allele was found in Patient 26 (rs199871047) and Patient 73 (rs139321356). The difference between genomic genotype (double peaks) and cDNA genotype (single peak) in Patient 26 and 73 implied the monoallelic expression of TAMM41. Red asterisks indicate the SNP positions. m Western blot analysis reveals that while a truncated protein caused by a premature stop codon was generated by the aberrant transcript identified in Patient 114, no protein was detected from the intron 6 inserted construct (Patient 26 and 73). n Co-immunofluorescence of mitochondrial outer membrane marker TOMM20 (red) and TAMM41-GFP showed that the truncated TAMM41, generated from Patient 114’s aberrant transcript, failed to localize onto mitochondria. Scale bar: 5 μm. V, vector. M, marker. White asterisks in e, f and j indicate transcripts generated by nest PCR. Red asterisks in f, g, j and k indicate the variation site
|

