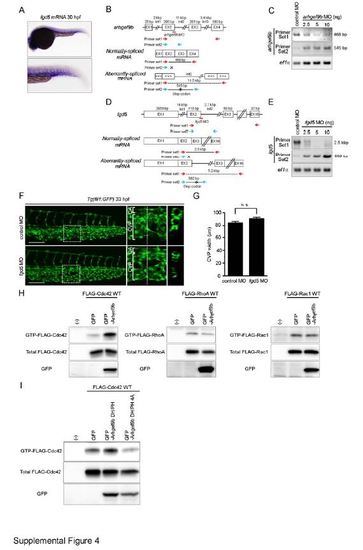
Arhgef9b, but not Fgd5, Is Involved in CVP Formation. Related to Figure 4. (A) Expression pattern of fgd5 mRNA in zebrafish embryo at 30 hpf, as detected by whole-mount in situ hybridization. The caudal region is enlarged at the lower panel. (B) arhgef9b MO (red line) is a splice-blocking MO that targets the boundary between exon 2 and intron 2. Schematic diagrams of arhgef9b gene (top), normally-spliced mRNA (middle) and aberrantly-spliced mRNA produced in arhgef9b MO-injected embryos (bottom). The size of exon/intron is indicated at the top of each exon/intron. EX, exon; Int, intron. (C) RT-PCR analyses of RNAs extracted from the zebrafish embryos injected with control MO or arhgef9b MO were performed using two sets of PCR primers as indicated in B (Primer set 1, red arrows; Primer set 2, blue arrows). PCR using Primer set 1 amplified the 468-bp fragment corresponding to normally-spliced arhgef9b mRNA in control MO-injected embryos, but not in those injected with arhgef9b MO. In contrast, PCR using Primer set 2 yielded the 545-bp fragment that contains partial sequence of exon 2 and intron 2 only in arhgef9b MO-injected embryos. These results suggest that arhgef9b MO blocks splicing of exon 2 and 3, leading to inclusion of intron 2 and the creation of premature stop codon indicated by asterisk. (D and E) Knockdown efficiency of fgd5 splice-blocking MO (red line) that targets the boundary between exon 2 and intron 2 were assessed by RT-PCR analyses similar to B and C. PCR using Primer set 1 (red arrows in D) yielded the 2.5-kbp fragment corresponding to normally-spliced fgd5 mRNA in control MO-injected embryos, but not in those injected with fgd5 MO. In contrast, the 982-bp fragment that contains partial sequence of exon 2 and intron 2 was amplified by PCR using Primer set 2 (blue arrows in D) only in fgd5 MO-injected embryos. These findings suggest that fgd5 MO blocks splicing of exon 2 and 3, leading to inclusion of intron 2 and the creation of premature stop codon indicated by asterisk. (F) Confocal z-stack images of the caudal regions of 33 hpf Tg(fli1:GFP) embryos injected with 2.5 ng control MO (upper panel) or 2.5 ng fgd5 MO (lower panel). The boxed areas in the left column are enlarged in the middle column. The cross-sectional images of the areas indicated by dotted lines on the enlarged images are shown in the right column.CA, caudal artery; CVP, caudal vein plexus. Scale bar: 50 µm. (G) The width of CVP as observed in F was quantified similar to Figure 1H (control MO [n=12], fgd5 MO [n=14]). Note that depletion of Fgd5 did not affect CVP formation. n.s., no significance. (H) 293T cells were transfected with the plasmid encoding GFP or that encoding N-terminally GFP-tagged Arhgef9b (GFP-Arhgef9b) together with vector expressing FLAG-tagged Rho GTPases (FLAG-Cdc42, right column; FLAG-RhoA, middle column; FLAG-Rac1, right column). GTP-bound Rho GTPases were collected as described in the Supplementary Experimental Procedures, and subjected to Western blot analysis with anti-FLAG antibody (upper panel). Aliquots of cell lysates were also subjected to Western blot analysis with anti-FLAG (middle panel) and anti-GFP (lower panel) antibodies. (I) 293T cells were transfected with the plasmid encoding GFP, that encoding N-terminally GFP-tagged DH and PH domains of Arhgef9b (GFP-Arhgef9b DH/PH) or that encoding its catalytically inactive mutant (GFP-Arhgef9b DH/PH 4A) together with the vector expressing FLAG-tagged Cdc42. Levels of total and GTP-bound Cdc42 were analyzed, as in H.
|

