- Title
-
Cloning and developmental expression of the DEC1 ortholog gene in zebrafish
- Authors
- Yao, J., Wang, L., Chen, L., Zhang, S., Zhao, Q., Jia, W., and Xue, J.
- Source
- Full text @ Gene Expr. Patterns
|
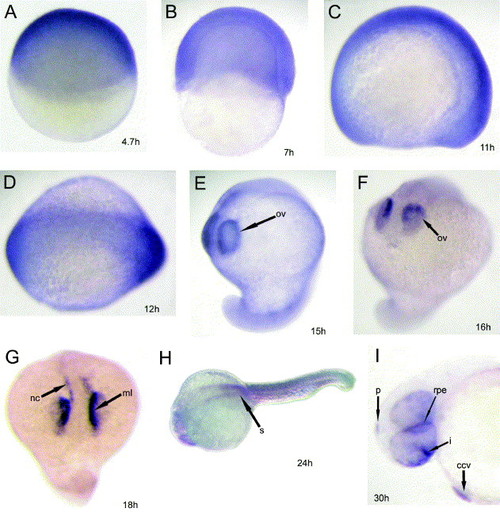
Expression of DEC1 during early zebrafish embryogenesis. All panels show a lateral view except panels (D, H, N and P) (dorsal view). Panels (O and U1) are cross sections cut through the planes indicated in panels (N (O)) and (T (U1)), respectively. DEC1 was strongly and widely expressed during early segmentation-30% epiboly stage (4.7 hpf, A), late shield stage (7 hpf, B), and 4-somite stage (11.3 hpf, C). The DEC1 signal declined at the 5-somite stage (12 hpf, D). Strong DEC1 expression was detected in the optic vesicle at 15–16 hpf (E and F) and in the medial layer of the eye and neural crest at 18 hpf (G). At the prim-5 stage (24 hpf, H), DEC1 was expressed weakly in the somite. From the prim-15 stage to the high-pec stage (30–42 hpf, I–Q), signals were detected in the pineal gland (epiphysis), iris, retinal pigment epithelium (rpe), common cardinal vein, blood cells and heart. (J′) A section cut through the heart. During this period, the DEC1 expression level in the somites appeared to be elevated (M–O and Q). At 36 hpf, signals were also detected in the notochord and pronephric duct (O), in addition, weak signals could be detected in the spinal cord (O). After 48 hpf, signals in somites decreased, but were still strong in the heart, rpe and pineal gland (R). At the pec-fin stage (60 hpf, S–U1/2), DEC1 expression in somites decreased, while weak expression in the hindbrain and hypothalamus began to appear. At this time, signals in the iris and pineal gland remained strong. U2, a section from a non-PTU treated control at corresponding developmental stage of U1 (60 hpf). At 65 hpf, DEC1 expression in the somites was no longer detectable, while expression in the notochord was obvious (V). At the protruding-mouth stage (72 hpf, W and X), strong expression in the pineal gland and in the retinal pigment epithelium (rpe) persisted, yet no obvious signals were found in the other regions. Abbreviations: bc-blood cell; ccv-common cardinal vein; h, heart; hb, hindbrain; ht, hypothalamus; i, iris; ml, medial layer; nc, neural crest; nch, notochord; ov, optic vesicle; p, pineal gland; pd, pronephric duct; rpe, retinal pigmented epithelium; s, somite; sc, spinal cord. EXPRESSION / LABELING:
|
|
Expression of DEC1 in early larva stages of zebrafish and in the adult brain. All panels show a lateral view except (F) (dorsal view) and G (ventral view). (C and D) Transverse sections through the midbrain and hindbrain of the 96 hpf embryos, as indicated in (B), respectively. Panels (F–H) are longitudinal sections through the swim bladder of 120 hpf embryos, with the anterior towards the left, and dorsal towards the top. (A) signals in midbrain and swim bladder at 84 hpf. (B–D) DEC1 expression is strong in the midbrain but weak in the hindbrain at 96 hpf. Meanwhile, the strong expression in the pineal gland remained and expression in the eyes was enhanced in the retinal. (E and H–J) The expression in the midbrain declined, but emerged in the swim bladder and digestive system including gut, liver and pancreas at 120 hpf. (F–G) DEC1 expression in the adult brain is mainly restricted in the anterior–lateral telencephalon, the most medial midbrain, and two lateral areas of the cerebellum, ventral diencephalons and myelencephalon. Abbreviations: dc, diencephalons; g, gut; hb, hindbrain; ht, hypothalamus; l, liver; mb, midbrain; myc, myelencephalon; p, pineal gland; pan, pancreas; r, retinal; sb, swim bladder; tc, telencephalon. EXPRESSION / LABELING:
|
Reprinted from Gene expression patterns : GEP, 6(8), Yao, J., Wang, L., Chen, L., Zhang, S., Zhao, Q., Jia, W., and Xue, J., Cloning and developmental expression of the DEC1 ortholog gene in zebrafish, 919-927, Copyright (2006) with permission from Elsevier. Full text @ Gene Expr. Patterns