- Title
-
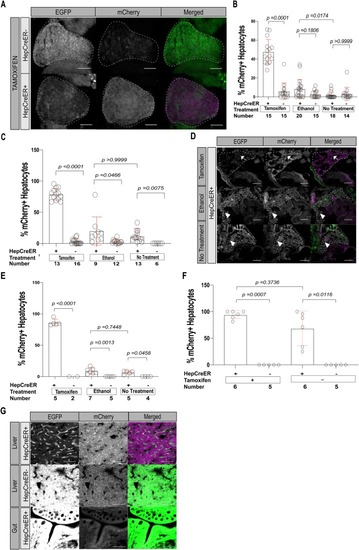
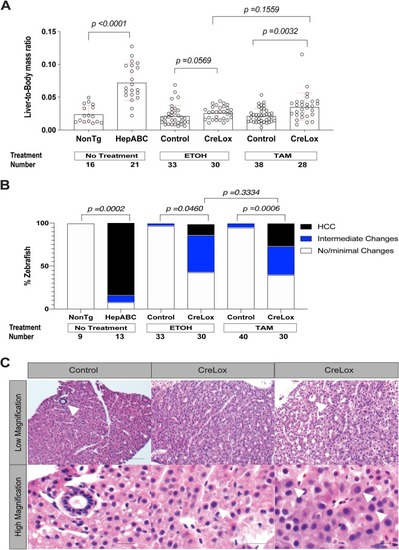
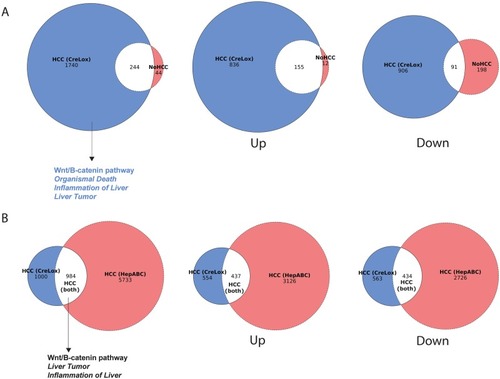
Heterogeneous beta-catenin activation is sufficient to cause hepatocellular carcinoma in zebrafish
- Authors
- Kalasekar, S.M., Kotiyal, S., Conley, C., Phan, C., Young, A., Evason, K.J.
- Source
- Full text @ Biol. Open
|
|
|
|
|
PHENOTYPE:
|
|
PHENOTYPE:
|
|
|
|
|
|
|