- Title
-
Evaluating Different Virulence Traits of Klebsiella pneumoniae Using Dictyostelium discoideum and Zebrafish Larvae as Host Models.
- Authors
- Marcoleta, A.E., Varas, M.A., Ortiz-Sever�n, J., V�squez, L., Berr�os-Past�n, C., Sabag, A.V., Ch�vez, F.P., Allende, M.L., Santiviago, C.A., Monasterio, O., Lagos, R.
- Source
- Full text @ Front Cell Infect Microbiol
|
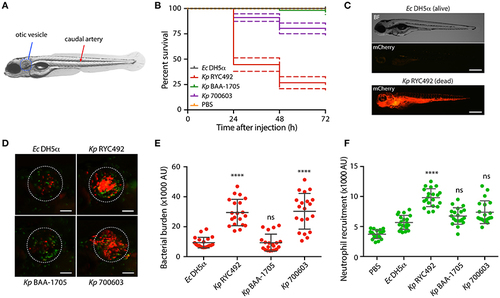
K. pneumoniae strains show different virulence behavior in the zebrafish larvae as revealed by survival curves and neutrophil recruitment. (A) Scheme of a 5-dpf zebrafish larva. Bacteria were injected in the dorsal caudal artery (red arrow) for establishing a rapid systemic infection, or in the otic vesicle (blue circle) for a localized infection. (B) Kaplan-Meier survival curves following injection of 3 dpf larvae with various strains of K. pneumoniae or with E. coli DH5? as a control. Larvae survival was monitored for 3 days after injection. The data are representative of three independent experiments, with a total of 57 larvae per bacterial strain. (C) Representative picture of a larva infected for 24 h with K. pneumoniae RYC492 strain, after acute bacteremia and death, and an unaffected larva injected with E. coli as a control. Bacteria expressing mCherry fluorescent protein is shown red. Scale bar, 500 ?m. (D) Neutrophils attracted to the otic vesicle of Tg(BACmpo:gfp) zebrafish larvae injected with K. pneumoniae strains or E. coli DH5? after 24 h post-injection (hpi). Bacteria are red and neutrophils are green. The white dashed lines encircle the otic vesicle. Scale bar, 100 ?m. (E) Bacterial burden in the otic vesicle of zebrafish larvae after 24 hpi. Measurements of red fluorescence from mCherry-labeled bacteria are plotted as arbitrary units (AU) for each K. pneumoniae strain or for E. coli DH5?. The data are representative of three replicates, with a total of 21 larvae per condition. (F) Neutrophils infiltrated in the otic vesicle of Tg(BACmpo:gfp) zebrafish larvae after 24 hpi. Measurements of green fluorescence from EGFP-labeled neutrophils are plotted as arbitrary units (AU) for larvae injected with K. pneumoniae strains, E. coli DH5?, or PBS. The data are representative of three independent experiments, with a total of 21 larvae per bacterial strain. Error bars represent the standard deviation among the replicates. Statistical differences compared to E. coli DH5? were assessed using a Kruskal-Wallis analysis followed by Dunn's multiple comparisons test. ****p < 0.0001. |
|
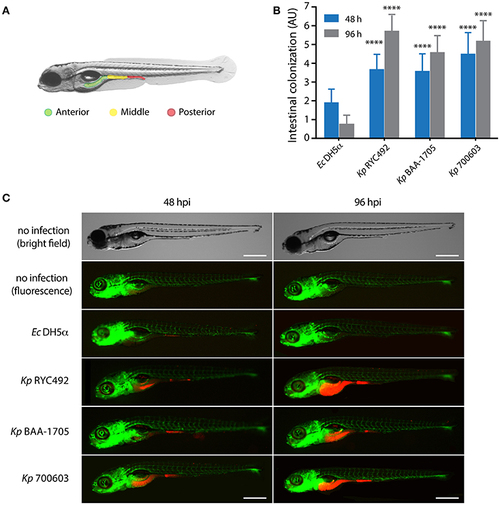
K. pneumoniae can infect zebrafish larvae by immersion and colonize its gastrointestinal tract. (A) Scheme of the gastrointestinal tract of a 5-dpf zebrafish larva, showing three arbitrarily defined zones (anterior, middle and posterior). (B) Semi-quantitative assessment of the intestinal colonization upon immersing zebrafish larvae in a suspension containing different K. pneumoniae strains or with E. coli DH5?. At the indicated time, larvae were anesthetized, observed under a fluorescence stereomicroscope, and the total intestinal colonization was scored in a scale from 1 to 9, following the criteria described in the Materials and Methods section. The data presented correspond to six replicates with a total of 75 larvae per bacterial strain. (C) Fluorescence microphotographs of zebrafish larvae representative of the data showed in (B). Bright field images of representative non-infected larvae at 48 and 96 h post-immersion are presented for more clarity. The larvae vasculature is shown in green, while infecting bacteria are shown in red. Scale bar, 500 ?m. ****p < 0.0001. |
|
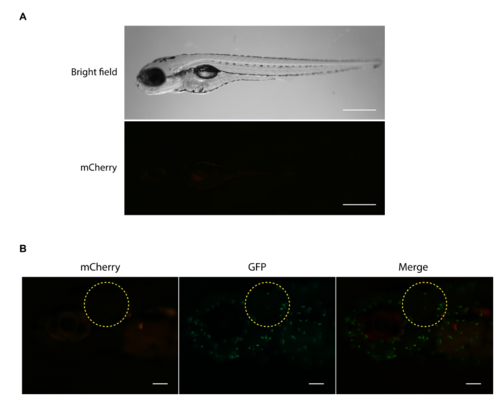
(A) Non-infected zebrafish larvae euthanatized by anesthesia overdose showed little or no red fluorescence, in contrast to larvae dead by bacteremia upon infection with Kp RYC492 (as shown in Figure 4). Scale bar, 500 ?m. (B) Larvae injected into the otic vesicle with sterile PBS (control condition) showed no red fluorescence in this area when evaluated 24 h post infection. The encircled area corresponds to the otic vesicle. Scale bar, 100 ?m. |