- Title
-
Comparative analysis of splice form-specific expression of LIM Kinases during zebrafish development
- Authors
- Ott, E.B., Te Velthuis, A.J., and Bagowski, C.P.
- Source
- Full text @ Gene Expr. Patterns
|
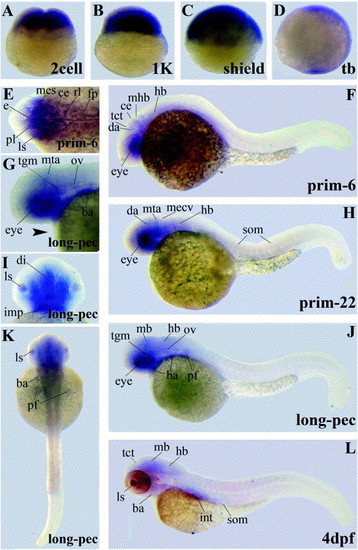
Gene expression patterns of LIMK1 variants encoding the PDZ domain. Abbreviations used are: branchial arches (ba), cerebellum (ce), diencephalon (di), dorsal aorta (da), epithalamus (e), floorplate (fp), hindbrain (hb), intermandibularis posterior (imp), intestine (int), lens (ls), midbrain (mb), midcerebral vein (mecv), mesencephalon (mes), midbrain–hindbrain-boundary (mhb), metencephalic artery (mta), otic vesicle (ov), pectoral fin buds (pf), proliferative cell layer (pl), rhombencephalon (rl), somites (som), tectum (tct), tegmentum (tgm). |
|
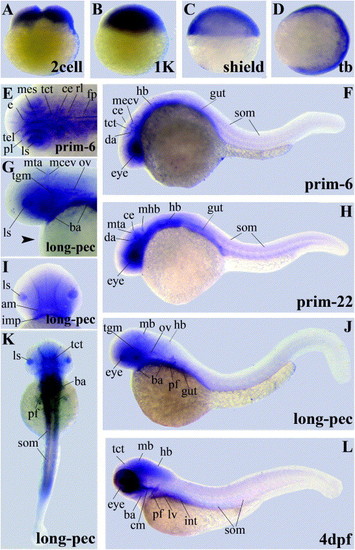
Gene expression patterns of LIMK1 variants encoding the LIM domains. Abbreviations used are: adductor mandibulae (am), branchial arches (ba), cerebellum (ce), cephalic musculature (cm), dorsal aorta (da), epithalamus (e), floorplate (fp), hindbrain (hb), intermandibularis posterior (imp), intestine (int), lens (ls), liver (lv), midbrain (mb), midcerebral vein (mecv), mesencephalon (mes), midbrain–hindbrain-boundary (mhb), metencephalic artery (mta), otic vesicle (ov), pectoral fin buds (pf), proliferative cell layer (pl), rhombencephalon (rl), somites (som), tectum (tct), tegmentum (tgm), telencephalon (tel). |
|
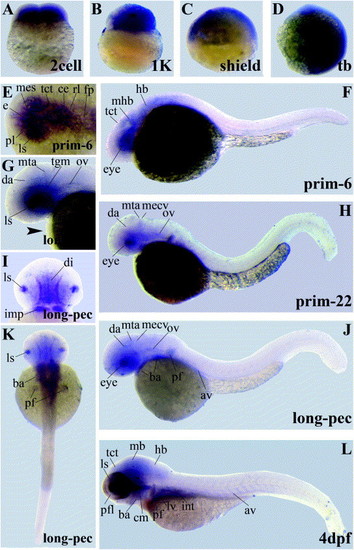
Gene expression patterns of LIMK2 variants encoding the PDZ domain. Abbreviations used are: axial vasculature (av), branchial arches (ba), cerebellum (ce), cephalic musculature (cm), diencephalon, dorsal aorta (da), epithalamus (e), floorplate (fp), hindbrain (hb), intermandibularis posterior (imp), intestine (int), lens (ls), liver (lv), midbrain (mb), midcerebral vein (mecv), mesencephalon (mes), midbrain–hindbrain-boundary (mhb), midcerebral vein (mecv), metencephalic artery (mta), otic vesicle (ov), pectoral fin buds (pf), proliferative cell layer (pl), rhombencephalon (rl), somites (som), tectum (tct), tegmentum (tgm). EXPRESSION / LABELING:
|
|
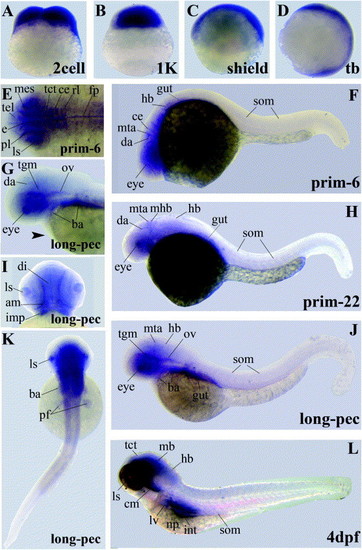
Gene expression patterns of LIMK2 variants encoding the LIM domains. Abbreviations used are: adductor mandibulae (am), branchial arches (ba), cerebellum (ce), cephalic musculature (cm), diencephalon (di), dorsal aorta (da), epithalamus (e), floorplate (fp), hindbrain (hb), intermandibularis posterior (imp), intestine (int), lens (ls), liver (lv), midbrain (mb), mesencephalon (mes), midbrain–hindbrain-boundary (mhb), metencephalic artery (mta), otic vesicle (ov), pancreas (p), pectoral fin buds (pf), proliferative cell layer (pl), rhombencephalon (rl), somites (som), tectum (tct), tegmentum (tgm), telencephalon (tel). EXPRESSION / LABELING:
|
|
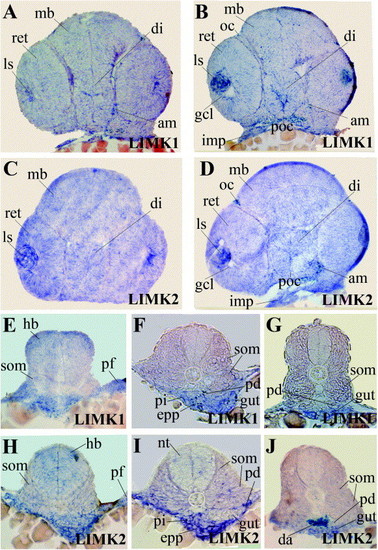
Shown are sections of whole mount in situ hybridizations of long pec stage embryos probed for LIMK1 and LIMK2, respectively. Abbreviations used are: adductor mandibulae (am), diencephalon (di), dorsal aorta (da), endoderm (e), exocrine pancreatic progenitor (epp), ganglion cell layer (gcl), hindbrain (hb), intermandibularis posterior (imp), lens (ls), midbrain (mb), notochord (nt), optic chiasm (oc), pectoral fin buds (pf), posterior chiasm (poc), pronephric duct (pd), somites (som), retina (ret). EXPRESSION / LABELING:
|
Reprinted from Gene expression patterns : GEP, 7(5), Ott, E.B., Te Velthuis, A.J., and Bagowski, C.P., Comparative analysis of splice form-specific expression of LIM Kinases during zebrafish development, 620-629, Copyright (2007) with permission from Elsevier. Full text @ Gene Expr. Patterns