Fig. 1
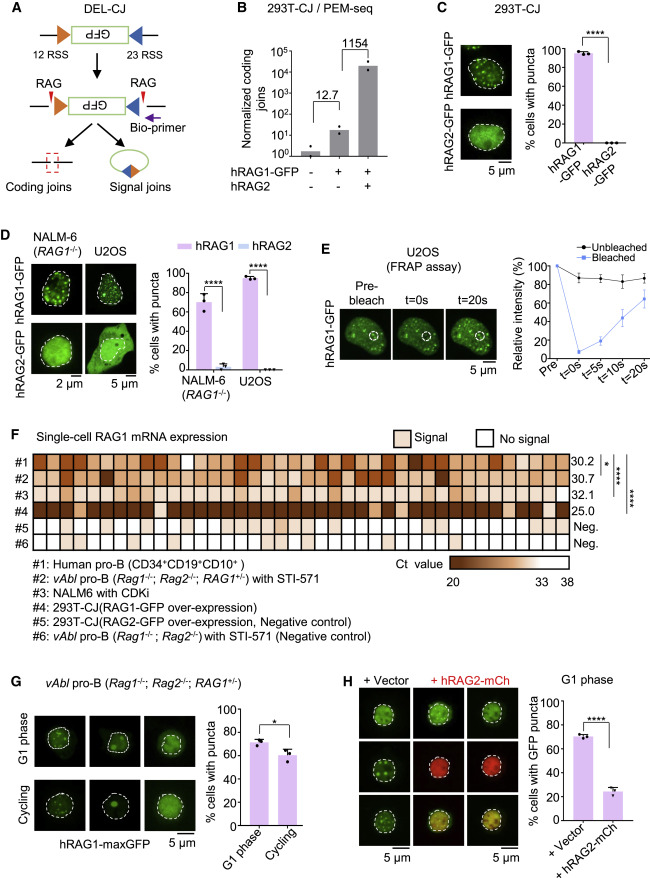
Figure 1. RAG1 undergoes aggregation in lymphocytes and nonlymphoid cells (A) Integration of bona fide 12 and 23 RSS pairs in HEK293T cells (293T-CJ), which can be recognized and cleaved by the RAG complex. Purple arrow indicates the position of the bio-primer for PEM-seq. Red triangles indicate the RAG complex. Green box indicates the inverted GFP segment. (B) Coding joins detected by the PEM-seq in 293T-CJ cells. Numbers of coding joins were normalized to total mapping reads including uncut and deletional coding joins. Fold changes are indicated. Replicates n = 2. (C) Representative microscopy images of indicated proteins in 293T-CJ cells (left). Bar graph (right) shows the percentage of cells with puncta in the total observed cells; replicates n = 3; t test; ∗∗∗∗p < 0.0001. Scale bar: 2 μm (left) and 5 μm (right). (D) Representative microscopy images of hRAG1-GFP in NALM-6 (RAG1−/−) and U2OS cells. Percentages on the right; replicates n = 3; t test; ∗∗∗∗p < 0.0001. Scale bar: 5 μm. (E) Representative images of hRAG1-GFP in U2OS cells for FRAP (fluorescence recovery after photobleaching) experiment. The dashed line indicates the circled bleached dots. Bleaching occurred at t = 0 s, and recovered intensity was detected at t = 20 s (left). Intensity recovery lines for hRAG1-GFP puncta are shown on the right. The background-subtracted fluorescence intensities are normalized by pre-bleach values. Cell number n = 15. Scale bar: 5 μm. (F) Heatmap showing RAG1 gene expression patterns across individual human pro-B cells or other types of cells in Figure S2D. The mRNA of RAG1 at single-cell level was quantified using the single-cell qPCR, and the Ct values were presented as color gradation. The average of Ct was marked for each sample. Cell number n = 40; t test; ∗p < 0.05; ∗∗∗∗p < 0.0001; Neg., negative control. (G) Representative microscopy images of hRAG1-maxGFP in vAbl (Rag1−/−; Rag2−/−; RAG1+/−) cells at cycling or G1 phase (left). Bar graph shows the percentage of cells with puncta (right); replicates n = 3; t test; ∗p < 0.05. Scale bar: 5 μm. (H) Representative microscopy images of hRAG1-maxGFP with overexpressed hRAG2-mCherry in vAbl pro-B (Rag1−/−; Rag2−/−; RAG1+/−) cells at G1 phase (left). Bar graph (right) shows the percentage of cells with puncta; replicates n = 3; t test; ∗∗∗∗p < 0.0001. Scale bar: 5 μm. All error bars represent mean ± SD. See also Figures S1 and S2.
Image
Figure Caption
Acknowledgments
This image is the copyrighted work of the attributed author or publisher, and
ZFIN has permission only to display this image to its users.
Additional permissions should be obtained from the applicable author or publisher of the image.
Full text @ Cell Rep.