Fig. 3
- ID
- ZDB-FIG-230413-24
- Publication
- Xiao et al., 2023 - Endothelial Brg1 fine-tunes Notch signaling during zebrafish heart regeneration
- Other Figures
- All Figure Page
- Back to All Figure Page
|
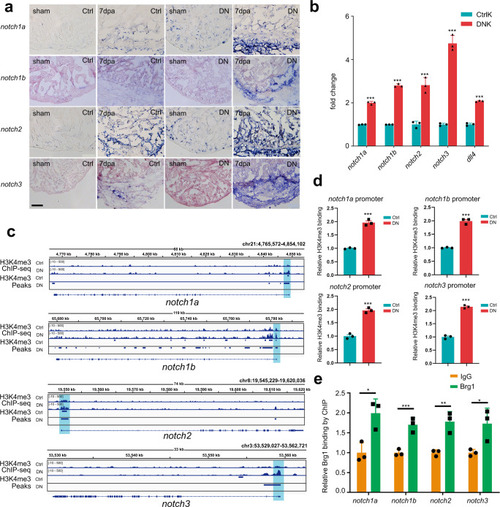
a Representative images of RNA in situ hybridization with notch1a, notch1b, notch2, and notch3 probes on frozen sections of sham-operated Ctrl hearts, injured Ctrl hearts, sham-operated DN hearts, and injured DN hearts at 7 dpa. Scale bar, 100 μm. b Quantitative RT-PCR analysis showing that the expression of notch receptors and ligand in FACS-sorted kdrl:EGFP endothelial cells from the DNK group was higher than those from the CtrlK group. Data represented one of three independent experiments. Data were mean fold changes ± s.e.m., ***p < 0.005, unpaired t-test. c H3K4me3 ChIP-seq showing the traces and peak intervals of representative genomic loci from Ctrl and DN hearts. H3K4me3 peaks in both Ctrl and DN groups were shown as bars. Putative promoter regions were indicated in blue color. d Anti-H3K4me3 ChIP and quantitative PCR in Ctrl and DN hearts at 7 dpa (primers designed from notch receptor genomic regions: notch1a, −171/+3 bp; notch1b, −41/+58 bp; notch2, −263/−115 bp; notch3, +394/+504 bp; ATG site designed as +1 bp). Data represented one of three independent experiments. Data were the mean fold changes ± s.e.m.; ***p < 0.005, unpaired t-test. e Anti-Brg1 ChIP and quantitative PCR in wild-type hearts at 3 dpa. Data represented one of two independent experiments. Data were the mean fold change ± s.e.m.; *p < 0.05, **p < 0.01, ***p < 0.005; unpaired t-test. n number shown here (b, d, e) indicated technical replicates. |