- Title
-
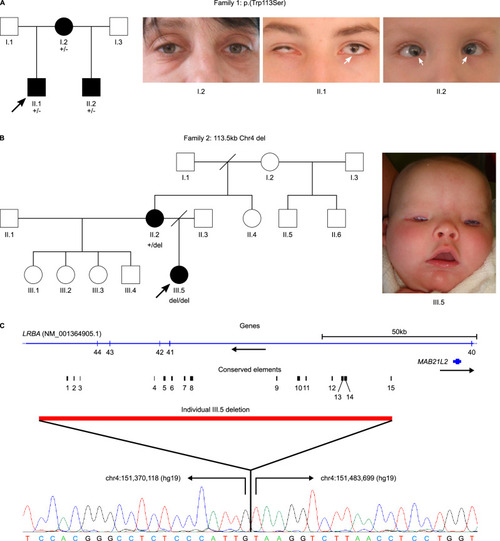
Deletion upstream of MAB21L2 highlights the importance of evolutionarily conserved non-coding sequences for eye development
- Authors
- Ceroni, F., Cicekdal, M.B., Holt, R., Sorokina, E., Chassaing, N., Clokie, S., Naert, T., Talbot, L.V., Muheisen, S., Bax, D.A., Kesim, Y., Kivuva, E.C., Vincent-Delorme, C., Lienkamp, S.S., Plaisancié, J., De Baere, E., Calvas, P., Vleminckx, K., Semina, E.V., Ragge, N.K.
- Source
- Full text @ Nat. Commun.
|
|
|
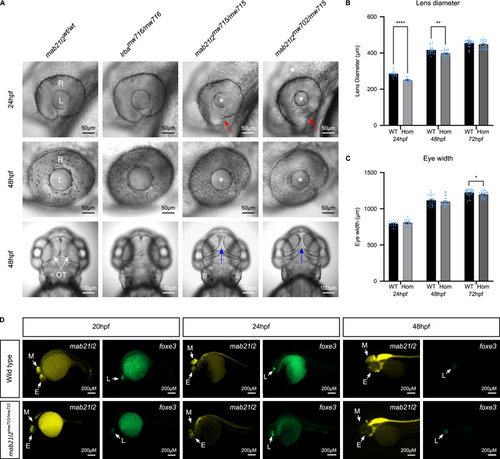
Zebrafish modelling of the chromosome 4 CNV showing disruption in eye and midbrain development. EXPRESSION / LABELING:
PHENOTYPE:
|
|
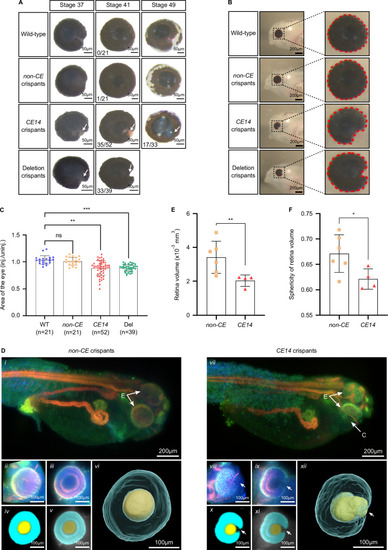
Characterisation of eye size and morphology in |
|
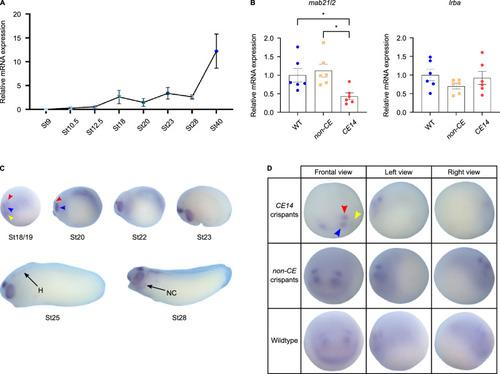
Expression studies of |

ZFIN is incorporating published figure images and captions as part of an ongoing project. Figures from some publications have not yet been curated, or are not available for display because of copyright restrictions. EXPRESSION / LABELING:
PHENOTYPE:
|